A CRISPR Approach to Immuno-Oncology
T cells can be tricky to engineer with CRISPR. Find out the key considerations when editing these cells and how you can overcome any associated challenges.
Join Us
Sign up for our feature-packed newsletter today to ensure you get the latest expert help and advice to level up your lab work.

T cells can be tricky to engineer with CRISPR. Find out the key considerations when editing these cells and how you can overcome any associated challenges.

Navigating the complexities of microbiome data visualization can be challenging. Our guide offers insights on selecting the right plots and optimizing them for maximum readability and publication readiness.

While CRISPR offers vast applications in disease research and drug target identification, it’s not always the optimal choice for every scenario. Explore the main advantages and challenges of using CRISPR-Cas9 to determine if it’s the right fit for your project.

Learn how the CRISPR prokaryote immune response systems were first discovered and the development of the CRISPR-Cas9 gene-editing tool.

Investigating human diseases and genetic variation is complex, but CRISPR-edited induced pluripotent stem cells present a promising alternative to immortalized cell lines. This article delves into genome editing principles and offers practical steps for optimizing research techniques, ensuring more accurate and ethical studies.

Genetic variants are critical to fields like evolution, diagnostics, and medicine—but they’re complex. This article breaks them down.

Maxam–Gilbert Sequencing. Slow and obsolete or niche but powerful? Discover how it works and learn about three modern applications.

Designing Cas13 gRNAs is a bit different from the standard Cas9. Read this guide to learn how it differs, and get a step-by-step guide on designing the perfect Cas13 gRNAs.

CRISPR isn’t just about DNA editing. Discover how you can use Cas13 proteins in your research to knock down, modify or track RNAs in mammalian cells.

This is the first installment in the DNA microarray series where I will introduce the technology and explain the basics.

If you need a multi-gene knockout or large-scale genomic modification, or want reduced off-target effects, then multiplex CRISPR is for you!

CRISPR interference allows for the regulation of gene expression in vivo. Here’s a short guide to how it works.

Discover how to check DNA quality for long-read sequencing using electrophoresis and why pipetting carefully is so important.

Discover how CRISPR can be scaled up for drug screening applications.

Find out how CRISPR-mediated gene activation (CRISPRa) and repression (CRISPRi) work and why you should consider using them in addition to your CRISPR knockouts.

Discover two CRISPR-based viral diagnostic strategies, DETECTR and SHERLOCK.

Find out how modified variants of CRISPR nucleases provide gene editing with reduced off-target effects and can even control gene expression without altering the DNA sequence.

Discover how to validate your CRISPR gene editing, from the successful delivery of CRISPR reagents to the confirmation of desired genetic and phenotype changes.

Discover how to get started with CRISPR gene editing in your experiments with our key considerations.

3′ mRNA-seq is a NGS method to profile gene expression that is sensitive and straightforward but won’t blow your budget.

Bioinformatics isn’t just for genomics geeks – there’s something for everyone!

Bioinformatics and NGS go together like peanut butter and jelly. But if you’re just starting out with these techniques it can be daunting.

The efficiency of whole genome sequencing (WGS) workflows has skyrocketed since its inception. Major leaps and minor tweaks in the WGS workflow have compounded over time resulting in radical reductions in processing time and the cost of sequencing whole genomes over the past decades. The complete sequencing of the first human genome, named the Human…

The success of whole genome sequencing (WGS) is shown in the quick and efficient scientific response to the 2011 outbreak of E. coli in Germany and France.1 German and French strains of E. coli were indistinguishable using standard tests. However, WGS analysis showed 2 single nucleotide polymorphisms (SNPs) in the German strains and 9 SNPs…

Confused about CRISPR nucleases? Read this guide to discover the various CRISPR nucleases available and what they are best suited for.

An oft-repeated maxim in biological bench science is that any experiment is only as good as its control. A control is an unchanging standard of comparison in an experiment, and an internal control is typically a standard reaction run together with the test reaction in the same reaction mixture. The purpose of an internal control…

Although multiplex CRISPR gene editing can be accomplished by simply introducing more than one gRNA to your target cells, there are many alternative — and more efficient — ways of achieving this goal. This article discusses these alternative CRISPR multiplexing strategies and highlights their potential caveats. Not sure whether multiplex CRISPR gene editing is right…

Want to do some epigenome editing? Discover the usefulness of catalytically inactive (dead) Cas9.

The advent of Next Gen Sequencing (NGS) has been truly amazing. One of the marvels that is often overlooked is how advances in DNA extraction technology have helped streamline NGS workflows. The original standard – phenol/chloroform extraction – is not well suited to the automated nature of today’s sequencing workflows (though with the emergence of…

Nanopore is a relatively new sequencing platform and researchers are still trying to optimize the protocol for their own specific applications. In our lab, we work primarily with metagenomic samples and use the 1D sequencing kits. Over the past year, we have optimized this technique. To check the quality of the Nanopore library preparation we…

CRISPRa allows you to activate or overexpress genes in a more endogenous manner. Find out the steps to getting started.
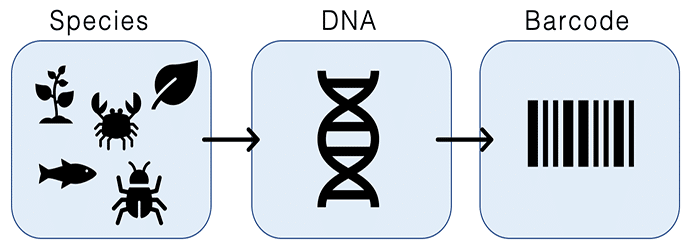
In both the lab and field, it is important to know what species we are working with. While morphological data has always been a tried and true method of identifying species, DNA barcoding allows us to identify species when we don’t have that option (e.g. if we don’t have enough of a specimen to identify…

Level-up your troubleshooting ability by determining the success of failure of each stage of your CRISPR experiment.

Reduced-representation genome sequencing has been one of the most important advances in the last several years for enabling massively parallel genotyping of organisms for which there is no reference-grade genome assembly. An implementation of the approach known as ddRAD-seq, first conceived in the Hoekstra lab at Harvard, has been widely adopted by the plant and…
WGS technologies have seen significant progress since the completion of the Human Genome Project in 2003. First-generation Sanger Sequencers were limited by lengthy run times, high expenses, and throughputs that read only tens of kilobases per run. The arrival of second-generation sequencers in the mid-2000s brought about the plummeting of sequencing costs and run times,…

In whole genome sequencing (WGS) initiatives it is not enough to simply sequence the whole length of the genomic DNA sample just once. This is because genomes are usually very large. The human genome, for example, contains approximately 3 billion base pairs. Although sequencing accuracy for individual bases is very high, when you consider large…

One of the most powerful methods of modern cellular biology is creating and analyzing RNA libraries via RNA-sequencing (RNA-seq). This technique, also called whole transcriptome shotgun sequencing, gives you a snapshot of the transcriptome in question, and can be used to examine alternatively spliced transcripts, post-transcriptional modifications, and changes in gene expression, amongst other applications….

Maybe you want to examine the entire transcriptome or maybe you want to investigate changes in expression from your favorite gene. You could do whole transcriptome sequencing or mRNA-seq. But which one is right for your project? From budget considerations to sample collection, let’s briefly look at both to see which might be best for your…
RNA-seq is based on next-generation sequencing (NGS) and allows for discovery, quantitation and profiling of RNA. The technique is quickly taking over a slightly older method of RNA microarrays to get a more complete picture of gene expression in a cell. Data generated by RNA-seq can illustrate variations in gene expression, identify single nucleotide polymorphisms…

Today, the gut microbiome is garnering a large amount of media attention for its role in human health and disease. From influencing immune responses to impact our brain, the gut microbiome is an important and necessary aspect of our life. So much so, that current investigations in the gut microbiome are focusing on developing biomarkers for…

The eBook with top tips from our Researcher community.