The genome-editing technology, CRISPR/Cas9, is well known for assisting researchers in manipulating the genome of target cells and has already been discussed in a few posts.
This article is about repurposing the typical genome editing technology to interfere with gene expression, known as CRISPR interference (CRISPRi). The purpose of CRISPRi is regulation of the genome instead of modification. Using this system you can either activate or repress gene expression.
From CRISPR to CRISPR Interference
The usual CRISPR system is composed of 2 components, a “guide” RNA (gRNA) and a CRISPR-associated endonuclease enzyme (e.g., Cas9).
In the CRISPRi system, you co-express a catalytically dead Cas 9 (dCas9), lacking endonuclease activity, with the gRNA. The gRNA is complementary to the region of the gene of interest you wish to repress or activate.
Put this article into practice
Choose a free resource to help you move forward
EBOOK
Gene Editing 101
DOWNLOAD
Blood Collection Tube Chart
Key Features of the CRISPR Interference System
1. Unlike the genome modifications induced by the typical nuclease active Cas9, the dCas9-mediated gene interference is reversible and does not permanently modify genomic DNA.
2. The dCas9 molecule retains the ability to bind to target DNA, based on the gRNA targeting sequence, but it cannot cleave the target DNA.
3. You can tag the dCas9 with transcriptional repressors or activators to regulate the expression of the ‘target gene’. The concept is schematically shown in Figure 1.
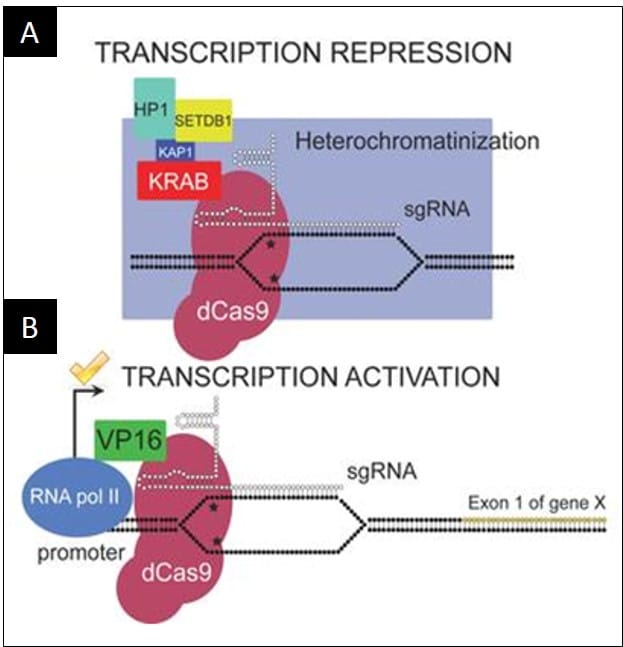
4. The simplest dCas9-based activators and repressors consist of dCas9 fused directly to a single transcriptional activator (e.g., dCas9-VP64 or dCas9-VP16) or transcriptional repressor (e.g. dCas9-KRAB).
5. The CRISPR-mediated gene activation, termed CRISPRa, is currently limited to gene activation in bacteria only.
How to Get Started with CRISPR Interference
Once you’ve decided on the desired genetic manipulation for your CRISPRi experiment (repression or activation), you need to select an expression system.
There are a few things you need to think about when you are selecting your expression system:
- Species of Cas9 and gRNA;
- Species of promoter and expression pattern of promoter for Cas9 and gRNA;
- Presence of a selectable marker (drug or fluorophore to quantify);
- Delivery method;
- Detection method.
After successful delivery of the gRNA and dCas9 (CRISPRi system) to your target cells, you will need to determine if your desired interference has occurred. Your detection method will depend on your specific application. Generally, after 3-5 days of transfection, you can dissociate CRISPRi cells and use flow cytometry to examine expression at the single-cell level. For more information, see our article on validating your CRISPR experiment.
The system can now handle up to five independent operations (activate/inhibit genes at 5 different sites) at once. However, the process will be limited by the number of guide RNA and proteins that can be inserted into a cell at a given time point, for the intended gene regulation.
The CRISPRi system goes beyond editing to repurpose the well-known CRISPR Cas9 system for precise genome regulation. The CRISPRi system holds great promise as a genetic programming platform, with potential applications in biomedical and clinical research.
Are you using CRISPRi in your research? Leave a comment with any helpful tips or tricks for getting it right below.
Originally published July 2016. Reviewed and updated May 2021.
Further Reading
- Dominguez AA et al. (2016) Beyond editing: repurposing CRISPR-Cas9 for precision genome regulation and interrogation. Nat Rev Mol Cell Biol. 17(1): 5-15.
- Gilbert LA et al. (2013) CRISPR-Mediated Modular RNA-Guided Regulation of Transcription in Eukaryotes. Cell. 154(2): 442-51.
- Larson MH et al. (2013) CRISPR interference (CRISPRi) for sequence-specific control of gene expression. Nat. Protocols. 8(11): 2180-96.
- Qi LS et al. (2013) Repurposing CRISPR as an RNA-Guided Platform for Sequence-Specific Control of Gene Expression. Cell. 152(5): 1173-83.
- CRISPR/Cas9 Guide
- Qi and Weissman Labs CRISPR plasmids
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
EBOOK
Gene Editing 101
DOWNLOAD
Blood Collection Tube Chart








