What Is a Cq (Ct) Value?
Don’t be confused by Ct values! We’ll guide you through what they are, how to calculate them, and troubleshooting issues.
Join Us
Sign up for our feature-packed newsletter today to ensure you get the latest expert help and advice to level up your lab work.

Don’t be confused by Ct values! We’ll guide you through what they are, how to calculate them, and troubleshooting issues.

Need to add extra nucleotides to your plasmid or other DNA sample? Here’s how to use overhang PCR to easily add extra bases using primers.

You can build your own PCR machine using tools and equipment you can get easily and without breaking the bank. Well, what are you waiting for?

Want to know the best way to perform relative quantification for your qPCR data? Read on.

Got non-specific PCR amplification? You need touchdown PCR! Discover what it is, how it works, and get 5 top tips for performing touchdown PCR.

Can’t be bothered doing a qPCR standard curve? Find out why it’s important to getting great results and how to do it.

Whether you’re generating, analyzing, or reviewing qPCR data you need to understand how it works and best practices. That’s why we’ve pulled together our top qPCR papers every researcher should read.

If you need to copy, sequence, or quantify DNA, you need to know about PCR. Read our guide to the PCR process, and discover tips to help you avoid the most common PCR pitfalls.

qPCR primer design is a bit of science, a bit of magic, and a little bit of luck. Here’s the science to help you design the best primers for your experiments.

New to qPCR? Here’s a quick summary of one of the two most common analysis methods – double delta Ct analysis.

As is sadly the case in many experiments, site-directed mutagenesis (SDM) does not always work the way we would like it to the first time around. Here are a few tips to help you on your way when trying to troubleshoot a bothersome SDM reaction!

How to Prevent False Results in Colony PCR Colony PCR saves time and reduces costs by eliminating the need for plasmid purification. However, confounding results abound — but only if you fail to anticipate them. This article outlines the major perpetrators of false results and how to prevent them. For a more general overview of…

Multiplex ligation-dependent probe amplification (MLPA) is a molecular technique developed by MRC-Holland back in 2002. In a nutshell, MLPA is a sensitive technique that allows quantification of nucleic acid sequences, quickly and efficiently. It is performed in many laboratories worldwide, and can be applied to detect copy number changes (like deletions or duplications) of a…
There are a few different ways of approaching site-directed mutagenesis. Here, I’ll give you a quick introduction to inverse PCR and why it’s useful, as well as going through a full protocol for SDM using modified primers!

Researchers have relied on immunodetection techniques such as Western blotting, flow cytometry and Enzyme-Linked Immunosorbent Assay (ELISA) for years, but immuno-PCR is a relatively new method. By merging an ELISA with the Polymerase Chain Reaction (PCR), immuno-PCR provides extremely high levels of assay sensitivity. ELISA An ELISA is an assay in which a molecule is…

The primers for your gene of interest have finally arrived in the mail, and you’re ready to figure out whether your favorite gene’s expression level is elevated in those precious tissue samples. Only one small last step before you can proceed. Primer validation. This is a standard procedure where you run PCR or qPCR on…

Every man, woman, and dog is doing quantitative real time reverse transcriptase PCR (qRT-rtPCR) these days. It’s a great method to measure your favorite transcript’s expression levels. One of the big plusses (like the Swiss flag!) of quantitative PCR in general is its high sensitivity. In principle, it can detect and quantify one molecule of…

Allele-specific expression can occur for various biological reasons, such as gene imprinting, or differential transcription caused by mutations, or single nucleotide polymorphisms (SNPs), or epigenetic alterations. Traditional end-point RT-PCR or qRT-PCR-based methods only detect overall levels of mRNA expression from a given gene rather than mRNA transcripts originating from individuals. If your project requires more…

Before we go any further, we have to get some things straightened out: RT-PCR versus qPCR versus RT-qPCR. Sooo confusing, amirite?? They all refer to specific molecular biology assays, but the names are unfortunately used interchangeably, which can be awfully confusing for just about anyone. So without further ado: Whew! I don’t know about you,…

I am sure many of you have been there. Everything is going smoothly, and your project seems to be working out perfectly. And then there is this one PCR. For some reason, it just won’t work. It is a black dot on your record. Even though I have a scientific mind, I have to be…

Here is a rundown of our top features to look out for when you are shopping for a new PCR instrument.

When you are a newbie to qPCR or qRT-PCR, it is quite common to write everything down in your lab notebook and then tediously mark off reagents or samples as you add them. People typically start with a master mix for items you add to multiple samples such as (Mg2+, dNTPs, 10X PCR buffer, additives,…
The unique feature of real-time quantitative polymerase chain reaction (RT-qPCR) is that it associates the amplification of your target gene with a fluorescent signal in a quantifiable manner. Presently, there are numerous fluorescent tool kits/methods to consider when designing your RT-qPCR experiment. However, the two major categories to choose from are fluorescent intercalating dyes and…

Who amongst us hasn’t had the need for oligonucleotides in an experiment? It is a cornerstone in many procedures and techniques. Depending on the goal, it can be very hard to design just the right oligo for your experiment. Oligos must have the right length; the right amount of C-G, T-A; they can’t form secondary…

It often happens that you do everything right with a PCR. You have perfectly isolated template DNA, used sterile tubes and tips, used clean reagents, and said a quick prayer to the PCR Gods. And still, something unknown messes up your results. This unknown at work is generally a PCR inhibitor. Before you blame it…

PCR was actually one of the first lab techniques I learned as an undergrad. Despite being sometimes labeled as a pretty basic lab skill, PCR doesn’t always work as expected. This “fickle” success is due to small details or hidden hazards within the PCR workflow that can cause your seemingly uncomplicated experiment to fail. This…

Droplet digital PCR? It’s easy. Because we’re here to guide you through it. We recently introduced you to the principles of digital PCR technology and how it differs from qPCR. In a nutshell, digital PCR is an end-point PCR technology that divides a single PCR into a large number of partitions, and then perform PCR…

Quantitative Reverse transcription PCR (RT-qPCR) is frequently used in the lab to detect and quantify RNA expression in a sample. The first step of the assay is to convert the labile RNA to its complementary DNA (cDNA) counterpart through reverse transcription (RT). In fact, RT is the first step in a variety of molecular biology…

There are many different methods and protocols on making your PCR run more efficiently. I recently came across an interesting PCR method called “nanoparticle” PCR. This method seems to attract a lot of attention, because it enhances a PCR by a few orders of magnitude. More interestingly, while the enhancement effect has been reported in a…
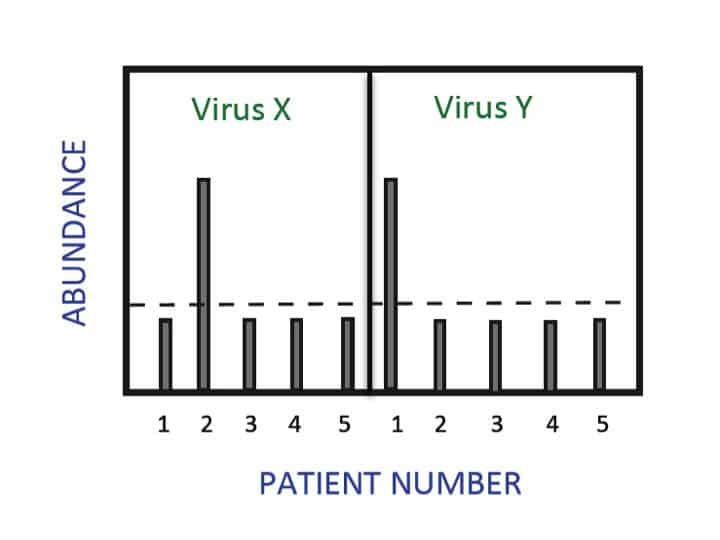
New ways to perform PCR emerge all the time. This speaks for the speed of technological advances, and reflects the ongoing need to keep up with fast-moving research. We all know that PCR’s main purpose is to amplify a stretch of nucleic acids based on sequence-specific primers. Nowadays, a wide range of PCR techniques exist,…

Good quality starting material is king for reverse transcription! Obtaining reliable results in any experiment requires good preparation. We often take reverse transcription for granted, and we don’t always consider that our qPCR might be performing poorly because of problems in that step. Since it’s quite often the reverse transcription reaction itself that causes fuss…
So, you’ve designed PCR primers to amplify your sequence of interest, and you’re ready to go. But unless you have a never-ending supply of template, polymerase, and a thermocycler with a gradient function—not to mention a hefty dose of time and patience—you probably don’t want to spend the next week finding the perfect conditions for…

DNA shuffling uses PCR technology in a very creative way. It allows you modify your protein to make a new protein you want. You can evolve proteins in microcentrifuge tubes on your very own lab bench. Isn’t that fantastic? DNA shuffling is also a very powerful technique for directed molecular evolution. W. Stemmer first used…

Kary Mullis invented polymerase chain reaction (PCR) in 1985 creating a revolution in molecular biology techniques. But it hasn’t stopped there. PCR has greatly evolved over the years. Today, we stand at a point, where we can clone micro RNAs (miRNAs) in real time! Due to miRNA size (about 18-21 nucleotides long) and varied expression levels,…

In the 30 odd years since its invention, the polymerase chain reaction (PCR) has become the bread and butter technique of molecular biologists. The secret to its indispensability lies in its simplicity and versatility. Numerous variants of the technique have been developed; one of these, real-time PCR, has become the method of choice for quantitative…
Routine PCR? Let’s be honest, there’s no such thing. Even with the simplest PCR reaction things can go wrong, so you need to have a good checklist of ideas for PCR troubleshooting and rectifying the problem. Today I have brainstormed all of the ways I can think of to approach problems with standard PCR reactions….

There’s piloting a brand new technique for the first time. Then, there’s jumping through hoops trying to get an established lab technique to work. The former, in contrast to the latter, is expected to be fraught with hardships. Yet troubleshooting an old lab technique that isn’t working anymore, is frustrating at a whole new level….

Do you work with plants? Are they genetically engineered? Do you know where and how? If not, you could experience problems. After all you do not want your transgenic gene cassette to disrupt genes that would affect your phenotype of interest. In this article I will tell you about Agrobacterium-mediated transformation – a widely used…

You’ve tried all the usual stuff, and checked the primer sequences twice, but still can’t get that PCR fragment amplified. It’s time to enter the strange world of PCR additives. Over the years a variety of additives have been shown to enhance PCR reactions in certain situations. Here is a summary of some of the…

The ability for DNA polymerase to copy a long stretch of DNA is becoming increasingly important. Why? It has to do with the advances in our sequencing technologies. Our next generation sequencing (NGS) technology requires the DNA polymerase to copy a long stretch of DNA (sometimes up to 50kb) as NGS is churning out genetic…

The eBook with top tips from our Researcher community.