It may not be mainstream (hipster hearts rejoice!), but imaging mass spectrometry (IMS) has been around for a while. IMS is a rapidly advancing technique that nicely blends the best bits of mass spectrometry with microscopic imaging. The result of which is a sort of ‘molecular histology’. Great for looking how drugs, lipids, metabolites, peptides, or proteins, are distributed in your sample (e.g. tissue of interest, tumor, biopsy etc).
How MS Works
If you are new to mass spectrometry, we’ve got some excellent primers How Does Mass Spec Work and Get Out of Western Blot Hell: An intro to Mass Spec. But in short, mass spectrometry (or mass spec to those in the ‘know’) allows you to measure the mass of molecules in a sample and quantify them. The same three basic steps in mass spec are in IMS too.
These Steps Are:
Ionization
Mass spec can only measure the mass of charged particles (i.e. ions). So, you must first break the molecules in your sample into smaller ions by blasting it with residual energy, using either a laser or bombarding it with electrons.
Mass filtration
Since opposites attract, positive ions created during ionization will move towards negative plates. The ions are then deflected by a magnetic field. The extent of the deflection depends on the mass, so ions of different mass will move through the machine at different speeds. Lighter molecules (i.e. those with a smaller mass) will move faster.
Detection
The final element of a mass spectrometer detects the charged particles one after the other as they hit the detector or fly past it. Those with a smaller mass are detected first. The result you get at the end of this process (which can be seen below in Figure 1) is called a mass spectrum!
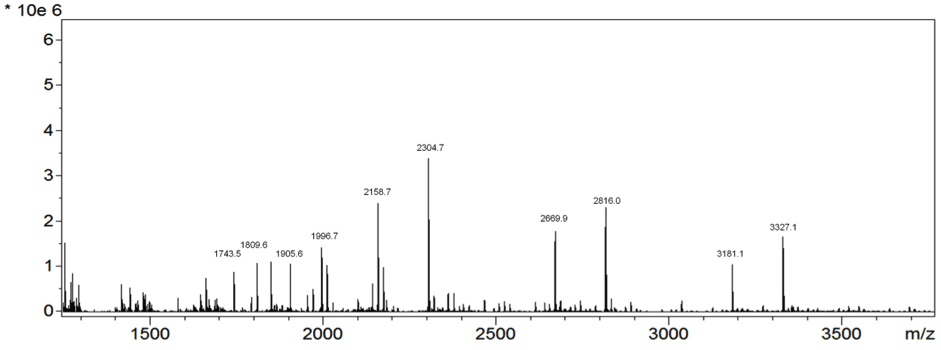
Figure 1. Adapted image of a MALDI spectra from mouse kidney tissue. Copyright (c) 2015 Thomas W. Powers et al. and made available on Wikimedia Commons under the Attribution- 4.0 International license
Enjoying this article? Get hard-won lab wisdom like this delivered to your inbox 3x a week.

Join over 65,000 fellow researchers saving time, reducing stress, and seeing their experiments succeed. Unsubscribe anytime.
Next issue goes out tomorrow; don’t miss it.
On the mass spectrum readout the mass-to-charge (m/z) values appears on the x-axis. And since the majority of the ions have a charge of 1, the m/z value is equivalent to the ion’s mass. The y-axis shows the intensity, which is a measure of the abundance of that ion in the sample.
How IMS Works
Usually to analyze your mass spec sample, you would perform the ionization, mass filtration, and detection, steps once – or if you are being stringent about replicability more times (3 seems to be the golden number!). The difference with IMS is that you scan your whole biological sample (e.g. a whole section of tissue), and sequentially collect a complete mass spectrum at each point. The distance between points is determined by the set scanning resolution.
Once you’ve collected your entire dataset then you can analyze one of two ways: 1) You can either look at a whole spectrum’s worth of ions expressed at a single point, or 2) Pick a certain m/z value and gauge its intensity at each point in your scanning raster. The latter approach is nice because it allows you to generate a ‘heatmap’ of your biomolecule of interest, visualizing its abundance throughout your region of interest.

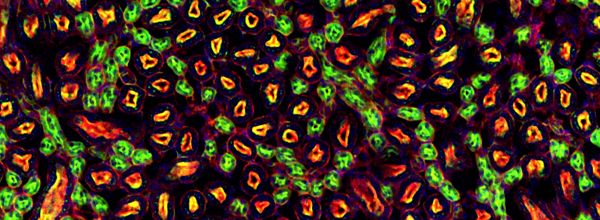
Figure 2. Adapted image of two ions that correspond to known N-glycans in the mouse kidney. The first Hex4dHex2HexNAc5 at m/z = 1996.7 is located in the cortex and medulla while Hex5dHex2HexNAc5 m/z = 2158.7 is more abundant in the cortex of the mouse kidney. An overlay image of these two masses is also shown. Copyright (c) 2015 Thomas W. Powers et al. and made available on Wikimedia Commons under the Attribution- 4.0 International license
The Magic of Heatmaps
If you are collecting spectra that are nanometers to micrometers apart, then these heatmaps are akin to microscopic images, but without the need for stains or antibody tags. This is why IMS is being termed ‘molecular histology’, see Figure 2. IMS is now increasingly being used alongside traditional histological and immunohistochemical methods, to provide information about a myriad of chemical compounds, be they drugs, lipids, metabolites, peptides or proteins, all in one run!
The whole IMS field was pioneered by Prof Richard Caprioli and his team, in the mid-90s, who used the MALDI (matrix-assisted laser desorption ionization) approach. However, MALDI isn’t the only IMS approach, SIMS (secondary ion mass spectrometry) is another emerging technology in the IMS field, and was introduced by Prof Nick Winograd and his team in the late-90s.
MALDI vs SIMS: What’s the Difference?
The main difference between MALDI and SIMS is the method of ionization. MALDI-IMS relies on an ultraviolet laser as a source of external energy to ionize the biomolecules in the matrix-coated tissue slice. In contrast to a laser, SIMS induces sample ionization using particle bombardment with a continuous beam of highly-focused, energetic ions such as Cs+, Au3 + and C60 +. The major analytical differences have been highlighted in the table below:
MALDI | SIMS | |
Spatial Resolution | Routine resolution: about 20µm-150 µm (depending on application) | Maximum resolution: order of 10nm |
Detectable Molecular Species | 100,000 species ranging from a few hundred Da up to 50 kDa | Generally larger than 1,000 Da |
Is IMS the right tool for you?
The main trade-off between the two IMS approaches is resolution (better in SIMS) and sensitivity (higher in MALDI). However, before you decide which IMS approach is the right one for your experiment, you must first determine if IMS is the right tool to answer your research question. IMS is the right tool if you want to visualize the distribution drugs, lipids, metabolites, peptides or proteins in your sample. But IMS also has limitations that you need to be aware of:
- It is still a good idea to stain your sample (or a sister-section) with hematoxylin and eosin to gain complementary histological information.
- IMS won’t work if you are studying a live biological sample, as the analysis occurs under high vacuum conditions.
- Biomolecules that are abundant in your sample are easier to detect. If your sample only contains minute amounts of your biomolecule, then you might have to take a different approach.
- IMS is great as a qualitative tool, but currently it can only be used semi-quantitatively. Absolute quantification of the same molecule in different samples, over different runs, can be difficult because there are other factors that can affect the ionization yield, and thus, the intensity recorded on the y-axis. Some of these factors include: 1) Slight variations in sample preparation. The laser-absorbing matrix is the key to MALDI’s success, but variations in its application (even within the same sample) can lead to intensity discrepancies. 2) Inherent technical variation due to the way the machine runs. 3) Anatomical differences. If your sample replicates are from different anatomical levels, your data might not be reproducible due to differences in the way your biomolecule of interest is distributed at those levels. However, you can use appropriate internal and external controls to account for these limitations.
- Finally, another major challenge of IMS, which many researchers grapple with, is the vast amount of data that these experiments generate. Processing IMS data often requires high computing power. Additionally, most of the software that is available for analysis is itself still at an experimental level.
Challenges, like those I’ve outlined above, are part and parcel of being a field in its infancy. Nonetheless, the future looks bright for IMS. It is definitely a powerful new tool that researchers can add to their analytical toolkit. Look out for more articles in this ongoing series to learn more about this new(ish) kick-ass technique!
———————–
Sources:
- Cornett DS, Reyzer ML, Chaurand P, Caprioli RM. MALDI Imaging Mass Spectrometry: Molecular snapshots of biochemical systems. Nature Methods 2007; 4: 828-833.
- Boxer SG, Kraft ML, Weber PK. Advances in Imaging Secondary Ion Mass Spectrometry for Biological Samples. Annu Rev Biophys 2009, 38: 53-74.
- Hanrieder J, Phan NTN, Kurczy ME, Ewing AG. Imaging Mass Spectrometry in Neuroscience. ACS Chem Neurosci 2013, 4(5): 666-679.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.