The first thing most of us do when entering the lab in the morning is play around with E. coli — defrost it, pellet it, suspend it, and so on.
It’s routine work that makes it easy to forget all the amazing uses of microbes in biotechnology.
Remember when we all made sourdough loaves during the COVID-19 lockdown? Biotechnology (sort of).
Love blue cheese? Biotechnology.
Put this article into practice
Choose a free resource to help you move forward
POSTER
Cell Culture Posters
POSTER
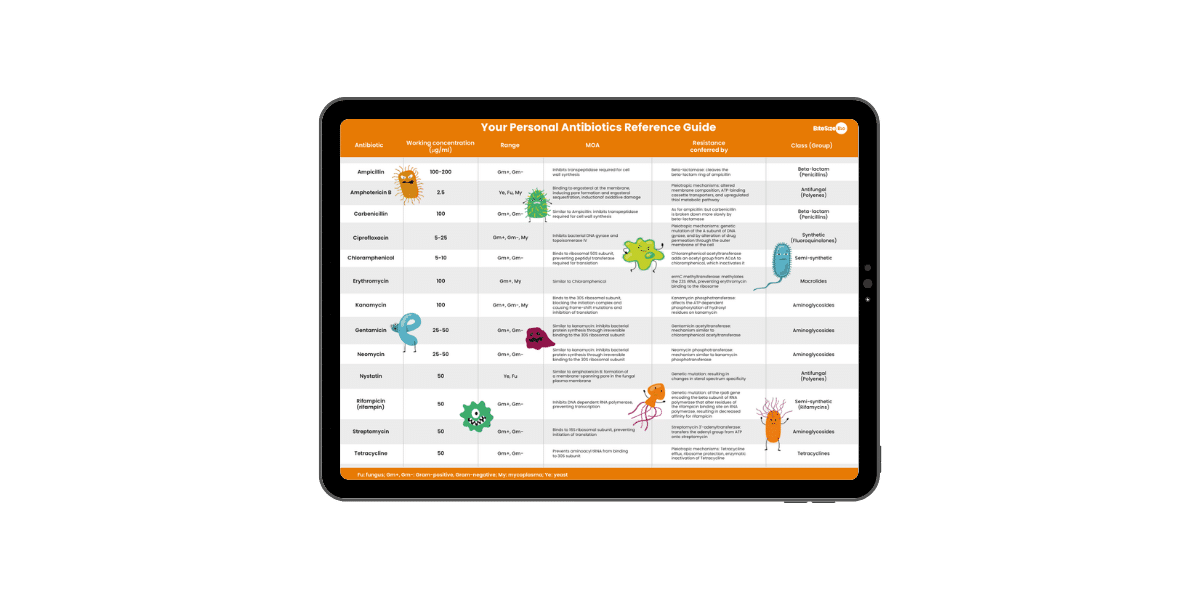
Antibiotics Reference Guide
Ever wondered how medical-grade insulin is made? We use bacteria to do it. Biotechnology.
If that’s piqued your interest, read on, as this article explains some of the more interesting ways we use microbes beyond lab research.
What is Biotechnology?
Biotechnology is the use of biological organisms, or parts of their cellular machinery, in technological processes.
It is almost as old as the civilization itself, although it wasn’t called “biotechnology” until the 20th century. Far from abandoning it in the 21st century, we continue to develop new uses for biological organisms.
Several journals focussing on biotechnology have emerged, such as Nature Biotechnology and Trends in Biotechnology, publishing articles from wide-ranging disciplines. This article will focus on our favorite uses of microbes in biotechnology.
1. Beer, Bread, and Wine
Biotechnology used for food and drink production is called yellow biotechnology.
Simple fungi in yeast forms were likely the first “domesticated” organisms used to make edible products through the process of fermentation. The consumption of glucose by yeast yields a by-product of ethanol and carbon dioxide, which can be exploited for generating bread, beer, and wine.
Likewise, the popular hipster drink, kombucha, and fermented cabbage (kimchi) are made through fermentation by yeast, lactic acid bacteria, and acetic acid bacteria. [1]
The most famous yeast is brewer’s and baker’s yeast Saccharomyces cerevisiae. It was first used by ancient Egyptians to make beer (even pyramid builders had beer rations). S. cerevisiae is harmless and easy to culture, and its culturing method was eventually refined by Louis Pasteur.
In 1996, the genome of baker’s yeast became the first sequenced eukaryotic genome. It is no wonder that it was the first eukaryotic organism used as a model for the production of various metabolites, from isopropanol to menthol.
Beyond yeast genetics, yeasts are used as a model to study protein-protein and protein-nucleic acid interactions, both of which have applications in drug discovery.
2. Milk Products
Fermenting milk using microorganisms is as old as domesticating herd animals, which began around 14,000 years ago in the Fertile Crescent region in Western Asia.
Milk is fermented by Lactobacillus; however, some cheeses, like blue cheese are made with Penicillium. Similarly, certain dessert wines are made from the “noble rot” of the fungus Botrytis cinerea.
3. Antibiotic Production
Red biotechnology is the use of biotechnology in the medical and pharmaceutical industries. It is called red because of the association with blood and symbols of medicine – red cross and crescent.
Fungi are most commonly associated with cheese and wine production. However, they have a long history in traditional medicine as moldy bread, overgrown with the fungi Penicillium and Aspergillus, would regularly be applied to wounds for their antimicrobial effect.
The jump from antimicrobial compounds to the antibiotics we know now was ushered in by Scottish scientist Alexander Fleming, who accidentally discovered Staphylococcus growth suppression by Penicillium mold in 1928.
However, it took another 12 years and a world war to start mass-producing penicillin using deep fermentation techniques. [2]
4. Restriction Enzymes
It had been known since the 1950s that certain bacteriophages caused poor growth of different bacterial strains, but the reason was unknown. In 1970, US researchers characterized an enzyme from the bacterium Haemophilus influenzae that can cut DNA in specific places.
This restriction enzyme, named HindIII, targets foreign DNA, protecting the bacteria from bacteriophages. [3]
In biotechnology, restriction enzymes are used to cut DNA to create a gap to insert a new piece of DNA. Today we know more than 3000 restriction enzymes recognizing 230 DNA sequences. [4] These restriction enzymes are used in biotechnology to cut DNA to create a gap to insert a new piece of DNA.
5. Protein Production in Bacteria
Protein production in Escherichia coli was a new stage in biotechnology because it used recombinant DNA technology instead of traditional selection techniques. The first human protein produced in E. coli in 1978 was insulin, [5] followed by human growth hormones. [6]
E. coli has become one of the most important microbes in biotechnology. Its genome and cellular machinery are well documented, making recombinant DNA techniques easy to design. Most importantly, it is extremely easy to culture using well-honed techniques that are routine procedures in most microbiology labs.
6. Microbial Fuel Cells
Scientists have worked out a way to harness the cell’s natural production of energy and direct it straight to an electrical circuit. These microbial fuel cells (MFC) are made using electroactive microbes like Geobacter and Shewanella, because they have specific properties that aid the transfer of electricity. [7]
There is enormous potential for MFCs to tackle sustainable energy production, where microbes can create electricity while simultaneously breaking down target substrates such as wastewater.
7. Phage Display Libraries
Similar to the discovery of restriction enzymes, the enzyme that joins the ends of recombinant DNA molecules, T4 ligase, also comes from a bacteriophage. But maybe the biggest phage contribution to biotechnology is phage display libraries (PDLs), developed in 1985. [8]
In PDLs, a gene encoding a protein of interest is inserted into a phage coat protein gene, causing the phage to “display” the protein on its outside. [9] PDLs are used in the studies of protein-protein, protein-peptide, and protein–DNA interactions. PDLs are useful in vaccine and drug discovery. [10]
8. Microalgae
Blue biotechnology uses sea resources to create products and industrial applications.
The leading application of blue biotechnology is producing renewable bio-oils with photosynthetic microalgae to replace oil extraction. Working with algae is not much different from working with other microorganisms.
To genetically modify your microalgae, you can use transformation, a familiar method of DNA introduction into a cell.
9. Agrobacterium Tumefaciens
Green biotechnology involves the genetic engineering of plants. It’s based on a neat trick by the microbe Agrobacterium tumefaciens. It has a plasmid, Ti, that transfers some of its genes into the plant genome. The transfer requires only T-DNA border sequences, so you can insert foreign DNA between the borders to be expressed in the plant cells. Several cereal crop plants were modified using this method, as well as HeLa cells. [11]
10. CRISPR
Last, but not least, in the history of biotechnology is the use of CRISPR. Of course, it’s not a microbe but rather a bacterial mechanism that is used to engineer organisms.
CRISPR was discovered in 1987 but has gained considerable traction in the last decade. In short, bacteria do not have a traditional immune system.
They evolved a mechanism to recognize bacteriophages that have invaded before. Each time a bacteria is infected with a bacteriophage, it stores a bit of DNA from the bacteriophage genome. Using the stored DNA sequence as a guide, the Cas9 enzyme (encoded by the bacteria) recognizes the invader DNA and destroys it.
Scientists recognized that this same mechanism could be used to introduce stable changes into the genetic material of eukaryotic cells as well.
Microbes in Biotechnology Summarized
Hopefully, having read this article and some of the others you will be able to see that the answer to the question “What have microbes ever done for us?” is “A LOT”! Microbes have contributed massively to the field of biotechnology with key roles in genetic engineering and the production of food and drink, therapeutics and vaccines, and biofuels.
References
- Villarreal-Soto, SA, et al. (2018). Understanding Kombucha Tea Fermentation: A Review: Understanding Kombucha tea fermentation. Journal of Food Science. 83(3): 580–588. doi:10.1111/1750-3841.14068
- American Chemical Society (2008). Penicillin Production through Deep-tank Fermentation. (Accessed 11/10/24)
- Smith HO, Wilcox KW (1970). A restriction enzyme from Hemophilus influenzae. I. Purification and general properties. Journal of Molecular Biology. 51 (2): 379–91. doi:10.1016/0022-2836(70)90149-X.
- Promega. Restriction Enzyme Resource. (Accessed 13/07/20)
- Human Insulin: Seizing the Golden Plasmid. Science News. 114 (12): 195. 1978-09-16. doi:10.2307/3963132.
- Goeddel DV, et al. (1979). Direct expression in Escherichia coli of a DNA sequence coding for human growth hormone. Nature. 281 (5732): 544–8. doi:10.1038/281544a0.
- Yee, M. O., et al., (2020). Cultivating electroactive microbes—from field to bench. Nanotechnology. 31 174003. doi:10.1088/1361-6528/ab6ab5
- Goff SP, Berg P (1976). Construction of hybrid viruses containing SV40 and lambda phage DNA segments and their propagation in cultured monkey cells. Cell. 9 (4 PT 2): 695–705. doi:10.1016/0092-8674(76)90133-1.
- Smith GP (1985). Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science. 228 (4705): 1315–7. doi:10.1126/science.4001944.
- Rahbarnia L. et al. (2017). Evolution of Phage Display Technology: From Discovery to Application. Review J Drug Target. 25 (3):216-224. doi: 10.1080/1061186X.2016.1258570.
- Kunik, T., et al. (2001). Genetic transformation of HeLa cells by Agrobacterium. PNAS. 98 (4): 1871–1876.. doi:10.1073/pnas.98.4.1871.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
POSTER
Cell Culture Posters
POSTER
Antibiotics Reference Guide