RNAseq libraries, also called whole transcriptome shotgun sequencing libraries, provide a snapshot of cellular processes. This allows the researcher to gain information regarding changes in transcriptome in response to environmental changes, during disease, or after a drug application. RNAseq libraries also allow for the detection of mRNA splicing variants and SNPs.
RNAseq libraries have virtually replaced microarrays because microarrays require a known template. Microarrays are also notoriously unreliable for detection and quantification of low and very high abundance RNAs and produce artefacts.
The exact procedure of constructing RNAseq libraries varies depending on the platform you are using. However, in general, all such libraries are obtained by the following steps:
1) RNA isolation,
Put this article into practice
Choose a free resource to help you move forward
PROTOCOL
Chemically Competent Cells Protocol
CHEAT SHEET
Nuclear Extraction Protocol
2) RNA fragmentation,
3) reverse transcription,
4) high-throughput (next generation) sequencing of cDNA
5) in silico sequence alignment1, 2.
This article is about the two first stages of making an RNAseq library (steps 1- 3).
Starting Material Preparation
As with any massive, multiple stage experiment, the quality of starting material is critical. For cell cultures, most of the cells should be in the same stage of growth. For tissue-specific gene expression, the tissue should be maximally purified from other tissue types.
The cells should be harvested quickly and with as little osmotic, temperature, etc. shock as possible. Flash freezing and grinding the resulting powder in liquid nitrogen is a preferred method to achieve minimally damaged nucleic acids.
If you are using RNAseq to assess a drug effect, preliminary experiments should determine an effective dose. If the drug concentration is too low, you won’t notice a transcriptional difference when compared to control samples; too high, and cells will shut down total mRNA production. For example, cycloheximide, the classic inhibitor of eukaryotic protein translation, has a IC50 of 532.5 nM and 2880 nM for protein synthesis and RNA synthesis in vivo, respectively.
RNA Isolation
As with any experiment involving RNA, you need to be extra careful to avoid RNA degradation. Autoclave all glassware in a high-temperature oven to reduce chances of RNase contamination. All buffers should contain RNAse inhibitors. Use certified RNAse free plastic and RNAse free DNAse. Be sure to decontaminate your workspace with RNase AWAY (or something similar), and always wear gloves.
Extract RNA from the tissues using your favorite method, for example, with trisol.
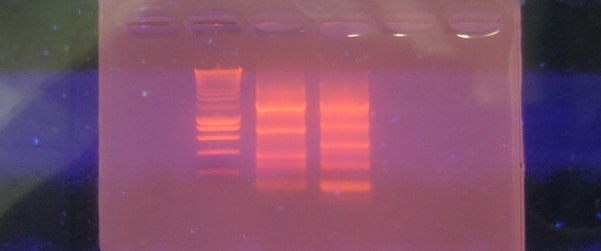
After RNA extraction, checking RNA integrity by electrophoresis, preferably capillary, is critical. RNA integrity number determined by RIN algorithm will tell you whether to proceed with the experiment or start again, as there’s no point in working with degraded RNA. The RIN ranges from 10 (intact) to 1 (completely degraded); you shoudn’t use samples with RIN below 73.
RNA Selection
If your sample passed the integrity test, you now have your total RNA. The next step is to get rid of types of RNA that you don’t want and enrich your sample with the RNA type you need. Start with removing rRNA as it constitutes 90% of your total RNA (really!) and will drown any signal. The simplest way to do this is to use a rRNA removal kit such as NEBNext rRNA Depletion Kit or Ribo-Zero rRNA Removal Kit by Illumina.
After eliminating rRNA if you are interested in total transcription levels, you can proceed to cDNA synthesis directly, otherwise you will need to further narrow down your targets.
Further RNA selection by type will require additional steps:
- To isolate mature mRNA, use the polyA tails that bind to poly(T) oligomers attached to beads. This method is suitable to assess mRNA splice variants.
- To isolate small RNA, such as miRNA, use size selection through gel filtration or affinity chromatography.
- Capture specific RNAs through hybridization with tailored probes.
cDNA Synthesis
DNA synthesis is probably the easiest step, if you RNA sample is high quality. Use 3’ or 5’ DNA linkers to hybridize with RNA to initiate reverse transcription. Treat with RNAse to remove the template.
Your cDNA is ready to sequence using a next-generation sequencing platform.
Although cDNA synthesis step is the most simple, it’s shown to introduce biases due to selective cDNA amplification4. However, if you are aware of this possibility, you can use corresponding controls.
In summary, the success of RNAseq libraries preparation depends on careful control at each stage. Happy sequencing.
Sources:
- Griffith M, Walker JR, Spies NC, Ainscough BJ, Griffith OL (2015). “Informatics for RNA Sequencing: A Web Resource for Analysis on the Cloud”. PLoS Computational Biology. 11 (8): e1004393. doi:1371/journal.pcbi.1004393.
- Wang Z, Gerstein M, Snyder M (2009). “RNA-Seq: a revolutionary tool for transcriptomics”. Nature Reviews Genetics. 10 (1): 57–63. doi:1038/nrg2484.
- Schroeder et al. (2006) The RIN: an RNA integrity number for assigning integrity values to RNA measurements. BMC Mol Biol 7:3. doi: 1186/1471-2199-7-3
- Liu D, Graber JH (February 2006). “Quantitative comparison of EST libraries requires compensation for systematic biases in cDNA generation”. BMC Bioinformatics. 7: 77. doi:1186/1471-2105-7-77.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
Nuclear Extraction Protocol
PROTOCOL
Chemically Competent Cells Protocol