Biosensor chip selection is a critical step in planning and running a surface plasmon resonance (SPR) experiment. Chip selection depends on the ligand or target that needs to be immobilized on the sensor chip, the analyte that is flowed over the target to study the binding, and the purpose of the biosensor assay (i.e., determination of kinetics, affinity, or analyte concentration).
In this article, we will discuss SPR chips, what they are made of, what surface coatings can be added and their uses, how to determine which type of chip to use for an experiment, and the latest advances in chip design and production.
The Sensor Chip Surface
A typical SPR biosensor chip is a glass chip coated with a thin layer of chemically inert metal, usually gold. This chip is normally functionalized by an additional chemical coating, called the immobilization matrix, which is attached to the chip via an adhesive linker layer (Figure 1).
Ligands or target molecules are attached to the immobilization matrix via either chemical coupling, where a permanent covalent bond is formed, or via capture coupling, where a non-covalent bond is formed.
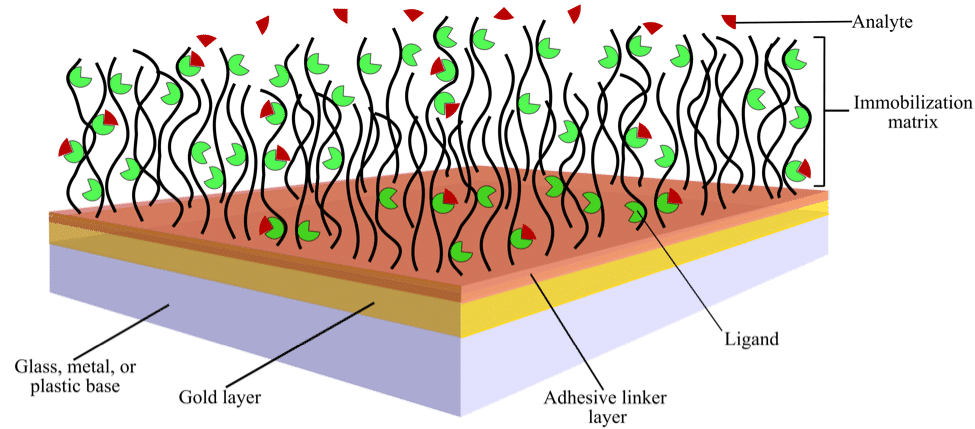
Figure 1. Schematic showing the surface of a typical sensor chip with immobilized ligand over which analyte is flowed
The Adhesive Linker Layer
The adhesive linker layer provides an anchor for the hydrophilic immobilization matrix and keeps it from being washed away (Figure 1). The thickness of this layer must be 2–5 nm because chip sensitivity declines significantly for layers greater than 10 nm in thickness. This is because the evanescent field strength decays exponentially with distance above the chip. Coatings thinner than the recommended range may be non-uniform and unstable.
Additionally, the refractive index (RI) of the adhesive layer must be lower than that of the base. Because SPR biosensing relies on the difference in RI between the sensor and the medium—a higher RI at the adhesive layer would interfere with detection. For chips coated with metals (for example, gold, as shown in Figure 1), alkyl derivatives of thiols, disulfides, and thioethers with hydrocarbon chains of length greater than 10 are commonly used as the adhesive layer. These form self-assembled monolayers (SAMs) on the metal surface with a high packing density. Bifunctional compounds, such as 16-hydroxy-hexadecane-1-thiol and 15-carboxy-pentadecane-1-thiol, are also used to form SAMs and can be used to directly couple target compounds.
For chips coated with glass or ceramic substrates (much less commonly used), the silanes, aminopropyltriethoxysilane and glycidylpropyltrimethoxysilane form monolayers to which either the immobilization matrix or the ligand can be attached.
Modification of plastic surfaces, another much less common substrate, involves dry or wet etching to generate surface layers carrying charged or polarizable groups. For instance, treatment of polyethylene with chromic acid or sulfuric acid generates surface layers carrying –COOH or –R2C=O groups. These surfaces are easily coated with appropriately charged polyelectrolytes. After etching, the plastic surface can be coated with a layer of stabilizing polymer or directly coupled to the immobilization matrix.
The Immobilization Matrix
The immobilization matrix minimizes nonspecific binding of the ligand to the adhesive linker layer. It often consists of a layer of hydrophilic polymers, such as dextran, alginic acid, pectin, carboxymethyl cellulose, carboxymethyl dextran, polyglycerol, polyethylene glycol, poly-L-lysine, or polyacrylic acid, among others.
These hydrophilic polymers form highly flexible, non-cross-linked, brush-like structures extending 100–200 nm from the surface (Figure 1). The three-dimensional nature of the hydrogels formed by these polymers offers the largest surface area for ligand binding of all matrix materials. Therefore, they are commonly used for studies involving low molecular weight analytes.
Planar or two-dimensional immobilization matrices are also available. These surfaces have a lower binding capacity and are suitable for studying interactions between proteins and other large molecules, such as high molecular weight proteins, viruses, whole cells, lipid monolayers, and lipid bilayers. Examples of such matrices include shorter chain dextran and SAMs of alkanethiolates.
Coupling of Ligands to the Biosensor Surface
There are four methods for coupling ligands to the biosensor surface—covalent, capture, ionic, and adsorptive.
Most Common:
- Covalent coupling involves formation of an actual chemical bond of the ligand to the sensor chip. A linker can be used to attach proteins via amide or other bonds.
- Ligands may also be affinity captured based on their specific binding to interaction partners bound to the immobilization matrix (Table 1).
Less Common:
- Ionic immobilization is used to couple highly charged ligands, such as oligonucleotides, to polyamine matrices. This method is rarely used, given its reversibility and instability with changes in pH, ionic strength, and the addition of chelating agents.
- Adsorptive methods are used for attaching ligands to plastic surfaces via hydrophobic interactions or gold or silver surfaces through thiol-metal bonds.
Below we discuss the two most widely used methods — covalent coupling- and affinity-based capture.
Covalently Coupled Biosensor Chips
Covalent immobilization of the ligand or target is carried out using the amine, thiol, aldehyde, carboxyl, or maleimide groups on the ligand.
Amine coupling is by far the most common choice because the main requirement of having available primary amines on the target is most easily met. A linker mix flowed over the immobilization matrix results in the addition of NHS esters that are then used to couple the target at free primary amine groups to form permanent ester bonds. Chips coated with carboxylated matrices, such as carboxymethyl dextran, are widely used for immobilizing ligands ranging from carbohydrates to proteins at high densities (Table 1).
The –SH groups of ligands can also be used for coupling via disulfide bond formation. This is less common because not all proteins have available thiol groups for coupling (although thiol groups can also be introduced). Disulfide bond coupling can be reversed by using reducing agents to regenerate the chip surface.
Carbohydrates and glycoproteins can be coupled by reactions involving their –CHO groups with hydrazide-derivatives of carboxylated matrices, but this is not commonly done.
Because covalent coupling involves forming a permanent covalent bond of the ligand to the sensor chip surface, only the analyte is removed after each sample injection (Table 2). An advantage of covalent coupling, particularly amine coupling, is that it is straightforward and points of multiple attachment are possible so high density surfaces can be prepared. Because a permanent bond is made, the surface is stable. However, ligand orientation cannot be controlled and may result in low or no binding of the analyte. Also, the ligand may be deactivated or denatured either during coupling or due to use of a regeneration solution between analyte injections (if needed) that is not suitable.
| Sensor chip surface chemistry | Applications |
| Long chain dextran/carboxymethyl dextran/alginate | General purpose; suitable for protein-protein interactions and small molecule analytes |
| Short chain dextran/carboxymethyl dextran/alginate or planar SAMs | Suitable for protein-protein interactions and for large analyte molecules and particles |
| Immobilized streptavidin | Capture of biotinylated ligands |
| Immobilized NTA | Capture of poly-histidine tagged ligands |
| Immobilized Protein A | Capture of IgG |
| Lipophilic modification | Capture of liposomes and supported lipid bilayers |
| Hydrophobic surface | Capture of lipid monolayers |
| Plain gold surface | Surface interaction studies and custom design of surface chemistry |
Table 1. Summary of biosensor chip surface chemistries and their applications
Capture Sensor Chips
Affinity capture relies on using a capturing molecule to attach the ligand to the surface of a sensor chip. To use this method, a capturing surface is first prepared either off-line or on-line. Ligand is then flowed over the surface and captured via an affinity-based approach. Examples of capture chips include the Ni-NTA chip for the capture of His-tagged proteins, a streptavidin or NeutrAvidin chip for biotinylated ligands, a protein A chip for IgG antibodies, hydrophobic chips for the capture of lipids, or antibodies for the capture of other antibodies or ligands (Table 1).
Ligand capture results in a temporary bond, except for Streptavidin/NeutrAvidin-biotin binding. The ligand is usually dissolved directly in running buffer when captured, so it is more likely to stay active. Most capture chips can be fully regenerated between sample injections, meaning both ligand and analyte are removed (Table 2), so ligand denaturation/deactivation is not common. However, this can also be a disadvantage. Fresh ligand is needed for each sample injection, which results in increased ligand or target consumption. Also, because the capture does not create a permanent bond, a decaying surface may result. Notable advantages of affinity capture are that ligand orientation is specific and highly purified samples of ligand are not required.
| Ligand immobilization method | Pros | Cons |
| Covalent coupling | Stable surfaceLower ligand consumption | Irreversible nature of ligand binding renders chip unusable if ligand is denatured or inactivatedRandom orientation of ligand, which can result in binding pocket being inaccessible |
| Amine-carboxyl or –CHO based | Easy to perform and consistentMost macromolecules contain –NH2 groups; therefore, amine coupling is the preferred method | Amine coupling not suitable for acidic ligands (pI < 3.5) |
| Disulfide-based | Binding is more specificLigand binding can be reversed using reducing agents | Some ligands don’t have available thiol groups (although they can be added)Ligand consumption can be high |
| Affinity capture | Ligand is specifically captured on chipDoes not require highly purified ligand In many cases, no permanent bond to the surface is made and the chip can be regenerated | Requires the added step ofcapture molecule immobilizationLigand consumption highIn many instances, ligand needs to be tagged |
| Biotin-avidin based | Given the very high affinity of biotin-avidin binding, ligand capture results in a stable surface | |
| Ni-NTA based | Convenient for the capture of ligands carrying poly-histidine purification tagsChip is easily regenerated using imidazole and/or EDTA | Can result in a decaying surface |
| Antibody based | Covalent coupling of antibodies provides a stable reusable chip | |
| Protein A-IgG based | IgG ligand is easily eluted using a number of methods, which include low pH, high pH, high ionic strength, and competitive binding, for chip regeneration |
Table 2. Pros and cons of the various methods for immobilizing ligands on the sensor ship surface
The Plain Gold Sensor Chip
The plain gold chip can be used to directly couple ligands containing natural or introduced thiol groups. Such an approach suffers from the problem of nonspecific binding of analyte molecules to the sensor chip owing to non-uniform ligand coverage, which may not be completely remedied by using blocking solutions. However, the SPR signal is enhanced by the proximity of the ligand and analyte to the gold surface.
A plain gold chip also provides researchers with an opportunity to coat the chip with new user-defined surface chemistries (Vashist et al., 2011, 2014).
In addition, it can be used to study redox reactions if an electrochemical accessory and a potentiostat are available for use with the SPR.
New Trends in Chip Design and Production
While a number of researchers have, over the past decade, tried to improve chip design, none of these newly designed chips has yet become widely used as a biosensor assay by the general SPR user population.
One such example is the use of metamaterials, especially those with negative refractive indices that amplify evanescent surface plasmon waves as well as convert them into propagating waves. These properties enhance the sensitivity of the chip to refractive index changes in the bulk medium. Proof-of-principle experiments using the biotin-streptavidin binding model have demonstrated the applicability of these materials to biosensing.
Substitutes for the glass base, the gold layer, and the adhesive layer have also been developed. Notably, an increase in sensitivity has been reported with the use of a ceramics base and a ZnO/Au nanocomposite instead of an Au/Cr layer.
Another trend is the integration of microfluidics with SPR chips. This generally involves fabricating microchannels on a suitable base, which is then bonded to the base bearing the SPR-compatible metal. The combination of microfluidics with SPR reduces the sample requirement and reaction time while enabling real-time label-free multiplexed detection within a single chip. This integrated technology promises to lower costs and is being explored for use in point-of-care diagnostics. It could also be valuable to researchers seeking automated high-throughput platforms.
No matter if you’re investigating protein-protein interactions, analyzing protein-small molecule binding, or conducting a more specialized assay, there is a sensor chip that can accommodate your application. The above review can help you make the decision about which sensor chip to use. For a comprehensive guide to SPR biosensor chip selection, check out our SPR chip selection guide.
Further Reading:
- Abbas A, Linman MJ, Cheng Q. (2011) New trends in instrumental design for surface plasmon resonance-based biosensors. Biosens Bioelectron. 26(5):1815–24.
- Arnoud Marquart. SPR Pages. Sensor Chips.
- Arnoud Marquart. SPR Pages. Immoblization.
- Chang CC, Chiu NF, Lin DS, Chu-Su Y, Liang YH, Lin CW. (2010) High-sensitivity detection of carbohydrate antigen 15-3 using a gold/zinc oxide thin film surface plasmon resonance-based biosensor. Anal Chem. 82(4):1207–12.
- Ekgasit S, Thammacharoen C, Knoll W. (2004) Surface plasmon resonance spectroscopy based on evanescent field treatment. Anal Chem. 76(3):561–8.
- Kravets VG, Schedin F, Jalil R, Britnell L, Gorbachev RV, Ansell D, et al. (2013) Singular phase nano-optics in plasmonic metamaterials for label-free single-molecule detection. Nat Mater. 12(4):304–9.
- Micheletto R, Hamamoto K, Fujii T, Kawakami Y. (2008) Tenfold improved sensitivity using high refractive-index substrates for surface plasmon sensing. Appl. Phys. Lett. 93:174104.
- Vashist SK, Dixit CK, MacCraith BD, O’Kennedy R. (2011) Effect of antibody immobilization strategies on the analytical performance of a surface plasmon resonance-based immunoassay. Analyst. 136:4431–36.
- Vashist SK, Schneiderc EM, Luongd JHT. (2014) Surface plasmon resonance-based immunoassay for human fetuin A. Analyst. 139:2237–42.








