Phenol extraction of DNA is a commonly used method for removing proteins from nucleic acids, e.g., to remove proteins from cell lysate during genomic DNA preparation. It’s commonly used, but not well understood.
If you want to know how phenol extraction works… read on.
DNA Extraction Using Phenol: The Basic Protocol
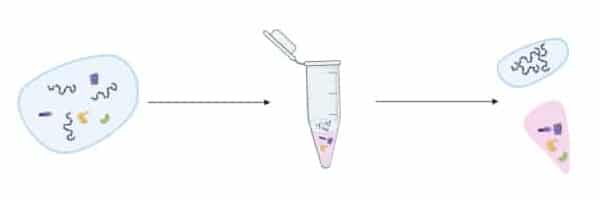
I’ll start with a quick outline of how phenol extraction of DNA is performed. First, a volume of phenol is added to the aqueous soup containing the proteins and the DNA to be purified.
Since phenol and water are immiscible (that is, does not form a homogeneous mixture when mixed), two phases form:
Enjoying this article? Get hard-won lab wisdom like this delivered to your inbox 3x a week.

Join over 65,000 fellow researchers saving time, reducing stress, and seeing their experiments succeed. Unsubscribe anytime.
Next issue goes out tomorrow; don’t miss it.
- a water phase (a.k.a. aqueous phase); and
- a phenol phase.
Phenol is the denser of the two liquids so it sits on the bottom.
The phases are then mixed thoroughly. This forces the phenol into the water layer where it forms an emulsion of droplets throughout. The proteins in the water phase are denatured and partition into the phenol, while the DNA stays in the water (Figure 1).
First, a Bit About Solvents
To explain how the addition of phenol can separate DNA and proteins, we need to briefly touch on solvents. This is the chemistry bit… bear with me.
A solvent is a substance, normally a liquid, that can dissolve other substances. Broadly, solvents can be classified according to their polarity, which depends on how extreme the spread of the electron density in the molecule is.
Water is a very polar solvent because the oxygen atom is very electronegative so it “sucks” the electrons towards it and away from the hydrogens, creating a slight negative charge on the oxygen and a slight positive on the hydrogens (Figure 2). That is, the charge is “polarised” within the molecule.
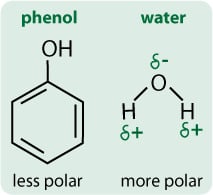
Phenol is a less polar molecule than water. Although it has a highly electronegative oxygen, this is counteracted by the phenyl ring, which is also very electronegative so there is no concentration of electron density around the oxygen. That is, the charge is not so polarised in a phenol molecule.
DNA is Most soluble in the Aqueous Phase
So what does this have to do with the separation of DNA and protein?
Well in general, polar (charged) compounds dissolve best in polar solvents and non-polar molecules dissolve best in less polar or non-polar solvents.
DNA is a polar molecule due to the negative charges on its phosphate backbone, so it is very soluble in water and less so in phenol. Therefore, when water (+DNA +protein) and phenol are mixed, the DNA does not dissolve in phenol. Instead, it remains in the aqueous phase.
Phenol Flips the Solubility of Proteins
But, proteins are a different story entirely. As you know, proteins consist of long chains of amino acids. Each amino acid has its specific characteristics owing to the nature of their side chains. Some, (e.g., phenylalanine, leucine, and tryptophan) are non-polar, because their side chains contain no charged entities. Conversely, amino acids with side chains containing charged entities (e.g., glutamate, lysine, and histidine) are polar.
The polarity differences in the side chains are biologically important because they largely determine how peptides fold into functional proteins.
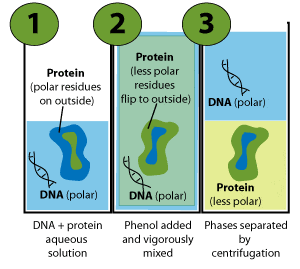
Put simply, the chains fold so that as many as possible of the side chains that are less polar than the solvent are inside the proteins (away from the solvent). In contrast, side chains of similar polarity to the solvent arrange outside (Figure 3, panel 1).
Another way to think about it is that polar side chains are hydrophilic, and non-polar are hydrophobic. The hydrophobic side chains hide on the inside, with the hydrophilic chains on the outside.
In the cell, cytoplasmic proteins fold according to the influence of water. However, protein folding changes upon exposure to a less polar solvent, like phenol (Figure 3, panel 2).
In short, the proteins flip inside-out, because the less-polar residues (usually hidden inside) now want to interact with the less-polar phenol.
Conversely, some of the very polar residues may flip to the inside of the globular protein to be shielded from the unsuitable new solvent.
In short, the proteins are permanently denatured by the new solvent environment provided by the phenol.
Whereas the polar residues on the outside of the proteins made them soluble in water, the phenol-induced folding changes forced the phenol-favoring residues to outside. The proteins are now more soluble in phenol than in the aqueous phase.
And this is the basis of phenol extraction of DNA. The phenol-soluble proteins partition to the phenol phase while, as discussed above, the water-soluble, polar DNA molecules stay in the water phase (Figure 3, panel 3).
Once you’ve obtained your aqueous DNA, you can use ethanol precipitation to extract it from the aqueous phase.
A Word of Caution
Phenol is a toxic chemical that is poisonous and corrosive. It can rapidly cause severe burns and should be handled with extreme care. (1) Because of the dangers of phenol, consideration of alternative methods should be given. Salting out is one method for DNA extraction that avoids the use of hazardous reagents like phenol. (2,3)
Final Thoughts
This article gives an overview of phenol extraction of DNA. Read our related article for a more in-depth look at the practical aspects of phenol/chloroform extraction, including differences between DNA and RNA extraction, the importance of pH, and why chloroform is used.
References
- Agency for Toxic Substances and Disease Registry. Medical Management Guidelines for Phenol. Accessed 14 October 2021.
- Miller SA, Dykes DD, Polesky HF. (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16(3):1215.
- Javadi A, Shamaei M, Mohammadi Ziazi L, et al. (2014) Qualification study of two genomic DNA extraction methods in different clinical samples. Tanaffos. 13(4):41-47.
Further reading
- Kirby, KS. (1957) A new method for the isolation of deoxyribonucleic acids: evidence on the nature of bonds between deoxyribonucleic acid and protein. Biochem J 66(3) 495–504.
- Pusztai A. (1966) Interactions of proteins with other polyelectrolytes in a two-phase system containing phenol and aqueous buffers at various pH values. Biochem J 99(1):93–101.
- Phenol extraction. Accessed 14 October 2021.
Originally published on February 12, 2008. Updated and revised October 2021.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.