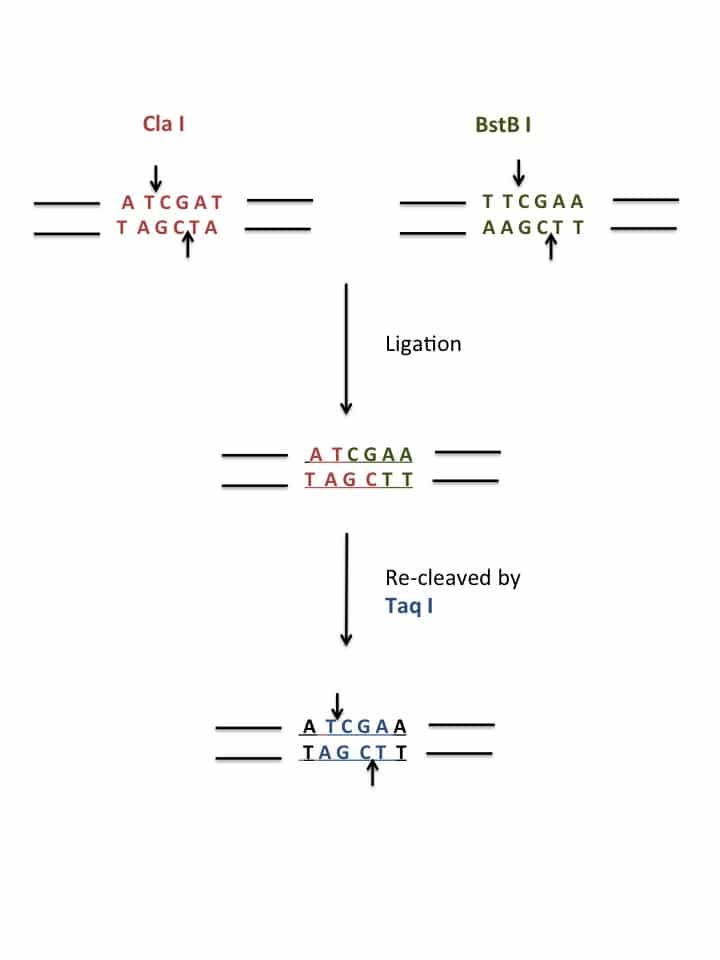
Restriction cloning, at its core, is quite simple. You simply cut the target vector and insert with the same enzymes, clean digested vector and insert up, ligate the two together, transform the ligated vector and insert into bacteria, and then screen. While getting each of the steps correct can be a bit of a hassle, in my mind, the most difficult part of the whole process is planning it.
If you are just starting with restriction cloning, it can be a bit of a struggle to learn the process, but that is ok. We are here to help! The easiest way to learn anything is to do it, so I thought I would take you step-by-step through the last cloning set-up I did. It had a little of everything, so it will cover most of the problems you will run into in your cloning endeavors.
Lay Out What You Have and What You Need for Restriction Cloning
Write down exactly which vector you have, what components you need where, and any other stipulations that are required at the end of the process. Make sure to make note of things like frame requirements, etc. This seems simple, but it is the most important part of the planning process.
For my cloning experiment I needed to remove the guide RNA sequences in a Cas9 vector that I had, and replace them with a new set of sequences I had synthesized by a company. I knew that I didn’t have to worry about anything being in frame, because that was already done for me. I also knew the new guides were being synthesized and shipped to me in a standard vector, pUC19.
Put this article into practice
Choose a free resource to help you move forward
PROTOCOL
Chemically Competent Cells Protocol
DOWNLOAD
Nanodrop Results Interpretation Card
Pull Up Your Target Vector, Desired Insert, and Other DNA Sequences on the Computer Screen
The first thing to do is open up your sequence files in your program of choice. I use SnapGene, but there are a large variety of programs out there that have these capabilities.
The first file to look at is the destination vector, so I opened mine up:
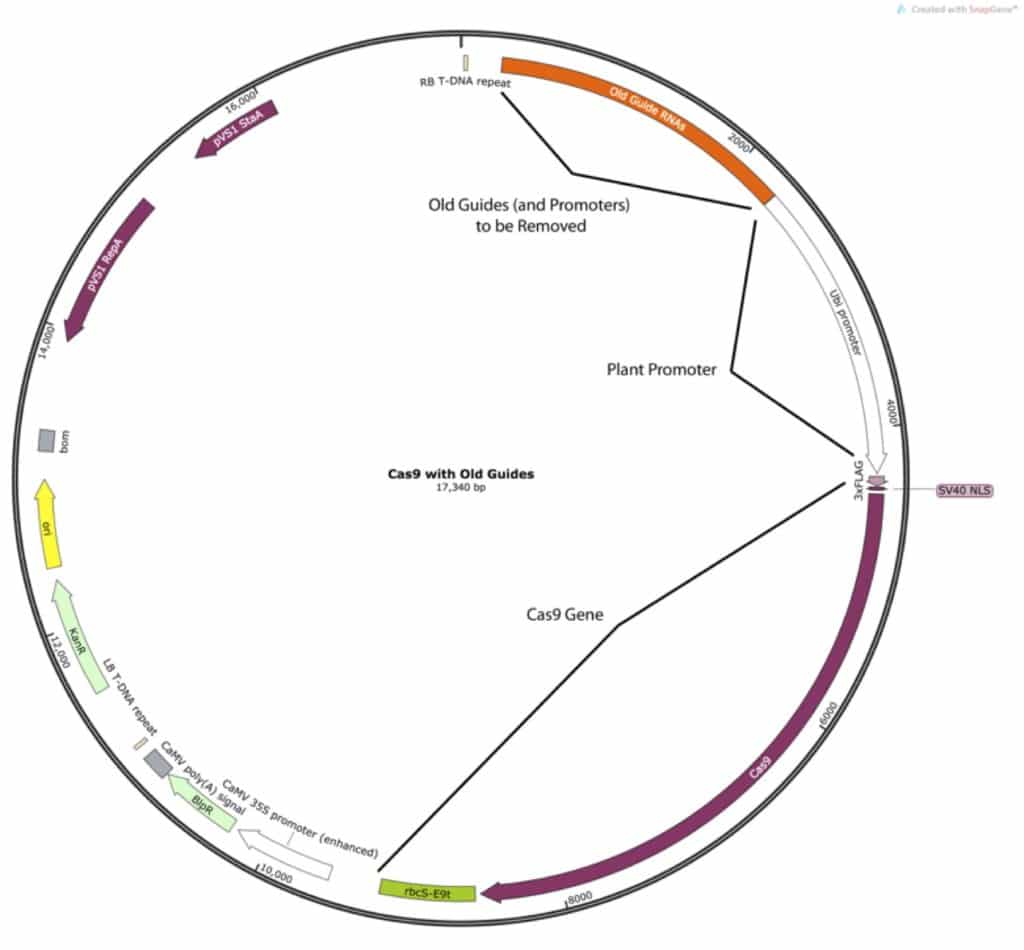
The vector consists of the old guides, a plant ubiquitin promoter to express the Cas9, a selection gene, and various plasmid maintenance genes. I was only concerned with removing the old guide RNA sequences and replacing them with those that I had designed:
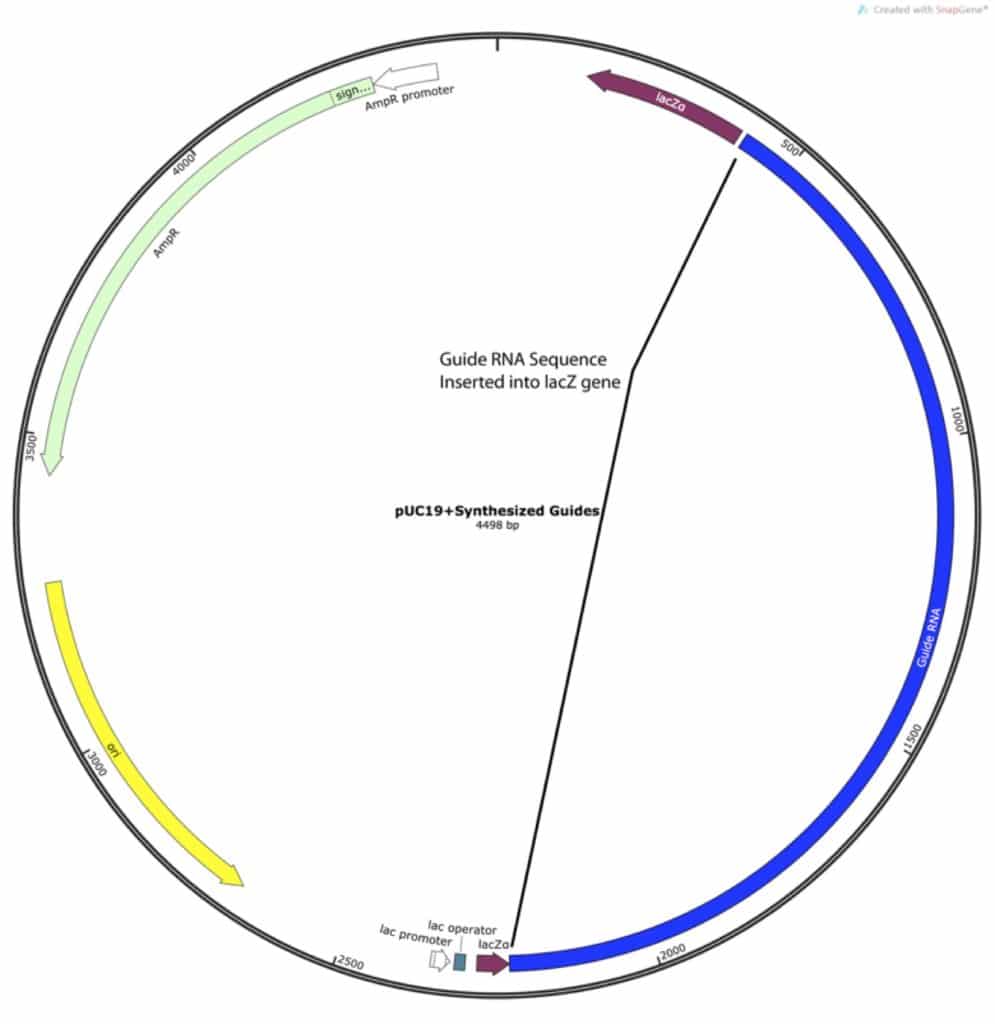
Find Compatible Restriction Sites
For restriction cloning to work, you need to find enzymes that cut both the target vector and the incoming DNA, so you need to examine the cut sites. For many cloning experiments, this can be complicated. It is important to consider how many times the enzyme cuts in the vector, the overhang that is left behind, and the type of enzyme recognition site. If at all possible, use enzymes that only cut once within a vector, leave sticky overhangs (that is 5′ or 3′ DNA overhangs), and have simple recognition sites (usually 6-8 base pair sites). Try and make the process as simple as possible, so that the actual cloning is easy as well. If you spend time carefully planning now, it will almost always pay off in the end.
For my particular cloning experiment I started by mapping the various enzyme sites on my vector.
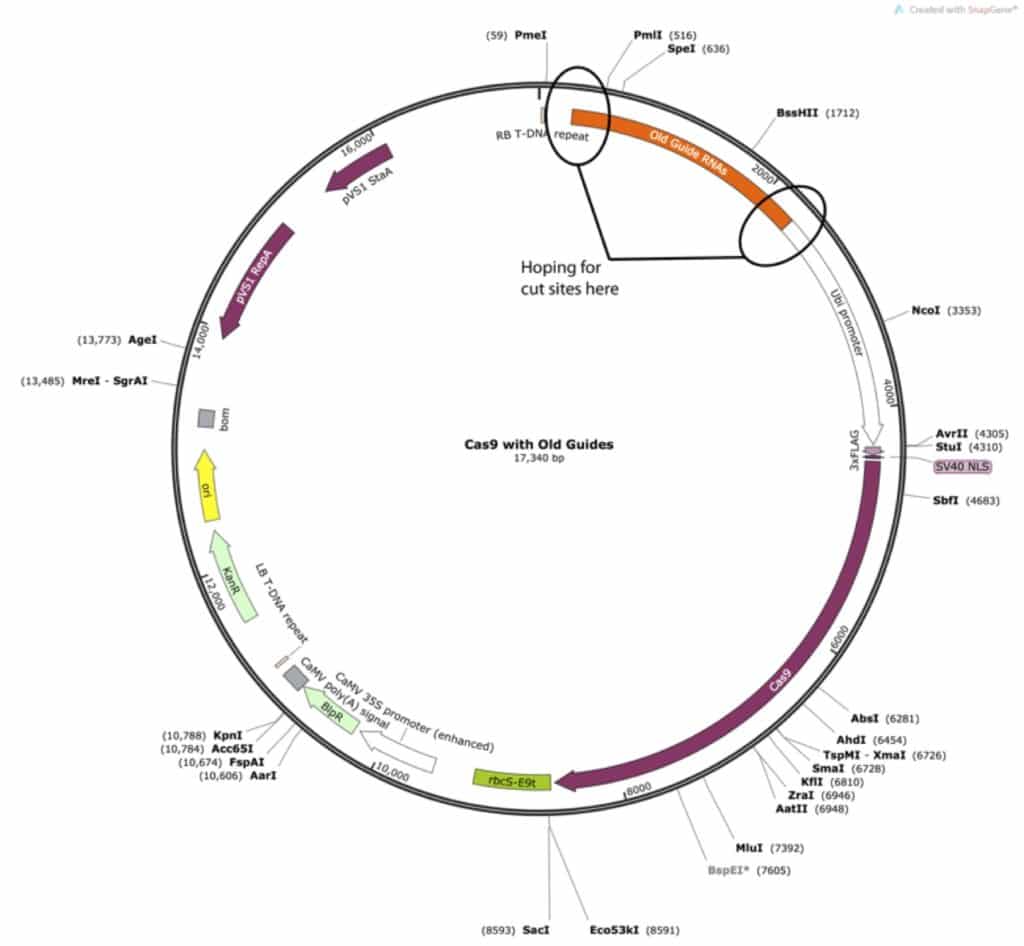
This is where I started to run into problems. As you can see, there aren’t any cut sites in between the old guide RNA and my promoter. If this sort of thing happens to you, don’t worry! There are almost always creative solutions to these sorts of problems. In my case, I noticed that while I couldn’t remove the guide sequences on their own, I could remove the guide and promoter. Since this is the common promoter used in my lab, I knew I could add it back in from another plasmid I had on hand.
Some other common problems and workarounds:
- There aren’t enzymes in common between the destination and insert: This is the most common issue you may run into.
- Consider enzyme overhangs: Some enzymes have overhangs that are in common and can be used together, despite not being the same recognition sequence, for example, BamHI and BglII.
- Consider filling in an overhang: While not ideal, a short treatment with T4 DNA polymerase or Klenow can fill in a sticky overhang and make it blunt. Blunt ends are always compatible, so can always work together. If you have to use a blunt site, try only to use it on one side of the cut, so that directionality is still conserved.
- Add cut sites with PCR: This option is best if you are already amplifying the insert via PCR. Simply add the recognition site to the end of the primer when ordering. Be sure to add some additional bases before the end of the primer to increase cutting efficiency. Once the PCR is finished, digest the product before ligation.
- The insert won’t be in frame
- If amplifying via PCR, add bases in the primer to ensure frame integrity.
- Consider cloning into an intermediate vector with different cut sites.
- Amplify the target with PCR using primers that add a cut site that is in frame.
- The enzyme cuts too many times
- This can be ok, depending on what you are working with. If the DNA is being removed from a plasmid anyway, as long as it doesn’t cut within the target, that is ok. Just make sure that you know what size bands you need to extract.
Work Around Any Issues
For my cloning experiment, I needed to replace a promoter with… the same promoter. Additionally, since one of the enzymes I would be using was PmeI, a blunt end cutting enzyme, I would need to make sure that I included a couple new sticky overhang restriction cut sites to use to insert my new guide RNAs to make the cloning easier. This is quite easy to do with PCR. I simply needed to find the promoter in a different vector:
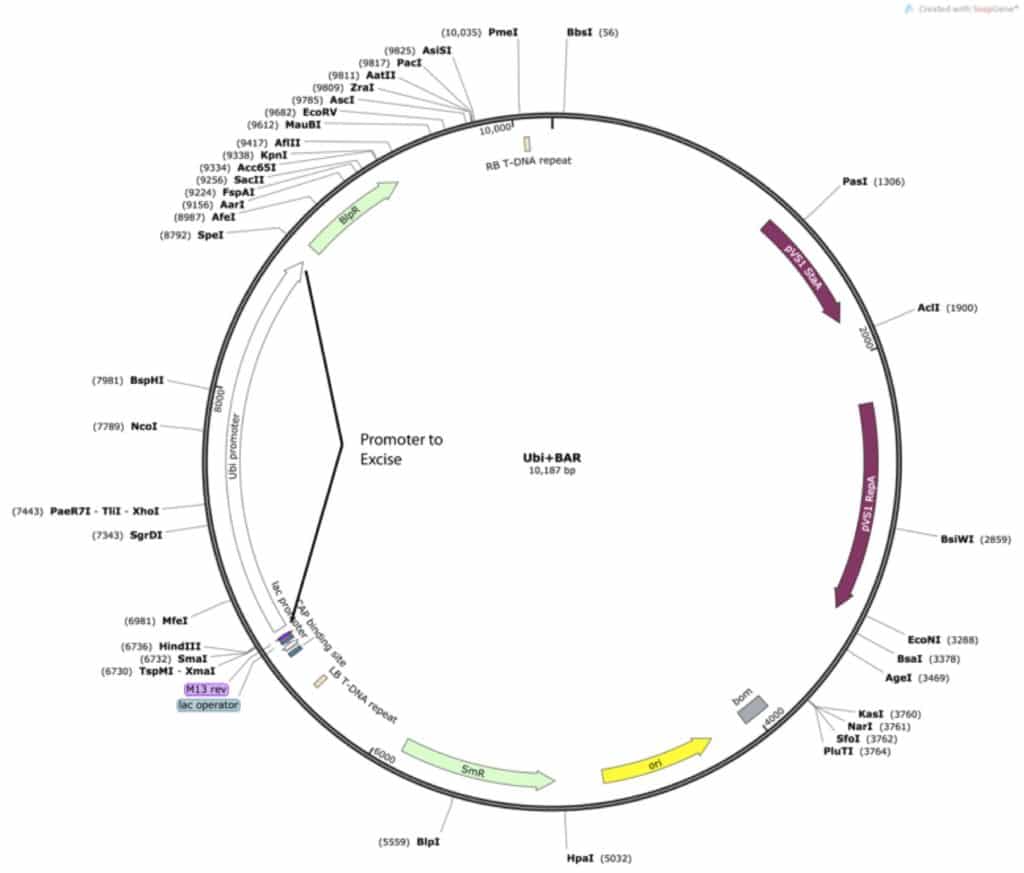
Unfortunately, none of the ones that I had would work perfectly with the cut sites I needed. I took care of this problem with PCR. I knew that my guide RNA sequences would most likely be delivered in pUC19, so I designed the PCR to include enzymes that would match those surrounding my new synthesized product. By designing primers with restriction sites attached, you can add the enzyme cut sites to the end of virtually any PCR product. Like so:
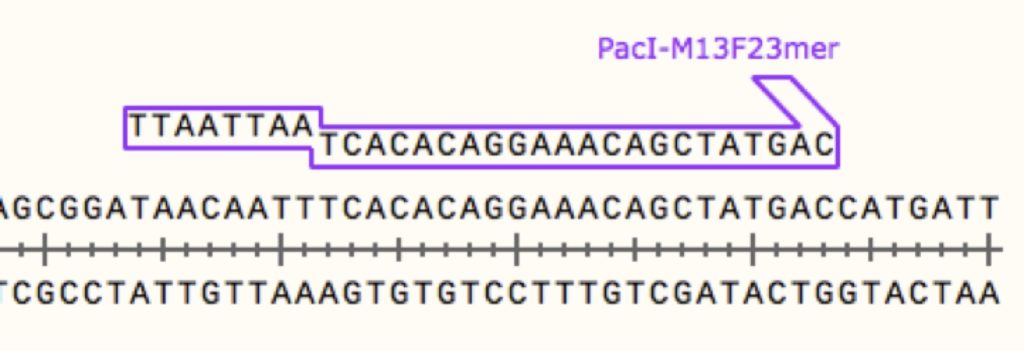
In this case the overhang of the primer will add the PacI restriction enzyme site, plus some additional bases for the enzyme to hold onto.
So for my cloning project I designed the primers to add PacI and AvrII to the end of my product:

After PCR I can digest the PCR product with AvrII, and the plasmid with PmeI and AvrII. One end of the PCR product (the uncut side) will then match the blunt end PmeI site, and one will match with the overhang left by AvrII. The cloning reaction result will be:
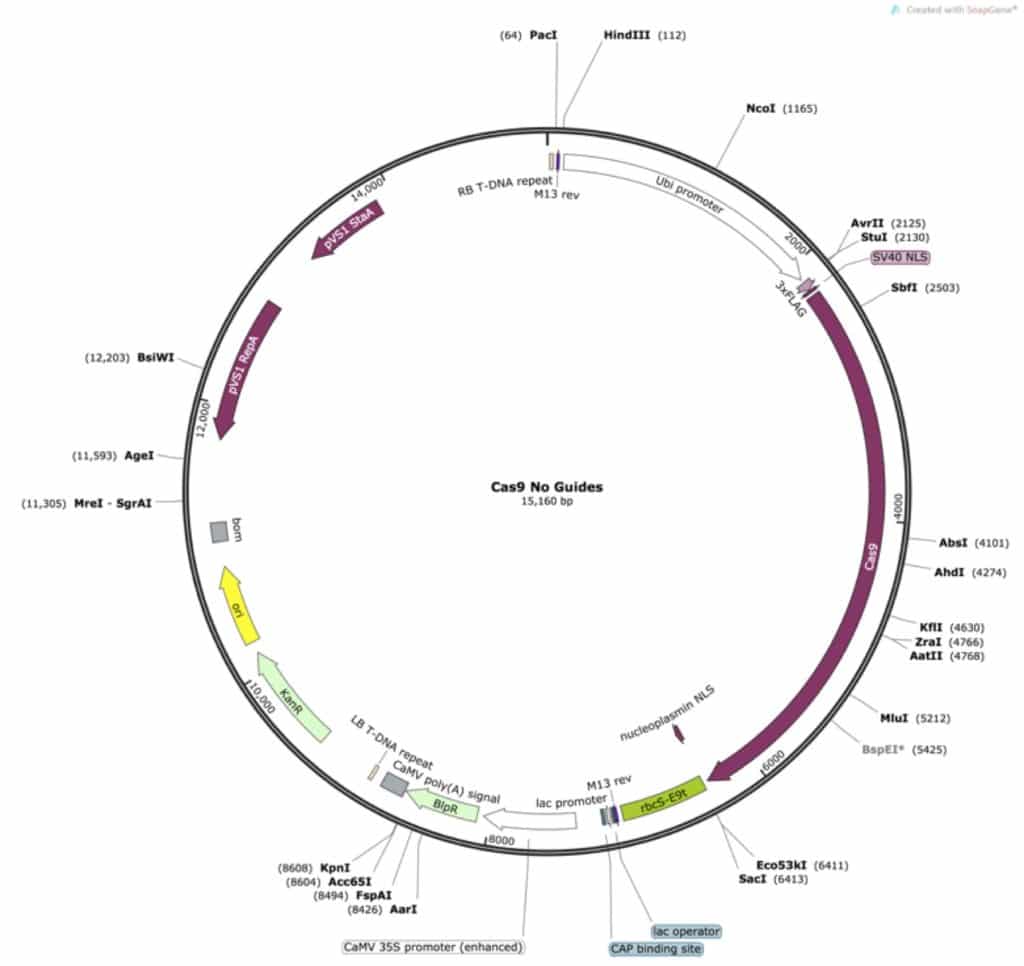
Perfect! My old guides are gone and my promoter has returned!
Check to Make Sure You Are on Track
At this point, return to your list you made at the beginning to make sure you haven’t forgotten anything important. It isn’t hard to lose track of things like frame when you are going back and forth between different cut site possibilities. Go back through your list and make sure everything is on track. Don’t be surprised if you have to tweak your plans a bit due to something unexpected happening. The good news is you can find these problems now, in silico, instead of having to figure them out while you are actually doing the cloning experiments.
Rinse and Repeat
At this point, you do the same steps as before ad nauseum until the entire experiment has been planned out. For my experiment I needed to now plan out how to insert the guide RNA’s into my new Cas9 vector. As you can see above, I only needed to flank the guides with PacI and HindIII to easily insert them before the ubiquitin promoter.
I was having my insert synthesized, and I knew that the company would insert the synthesized DNA into a vector to ship it to me. As a result, I saved a few bucks by finding an intermediate vector for them to insert the synthesized DNA into that would already have my required sites. I tried a few different options before I found one that worked.
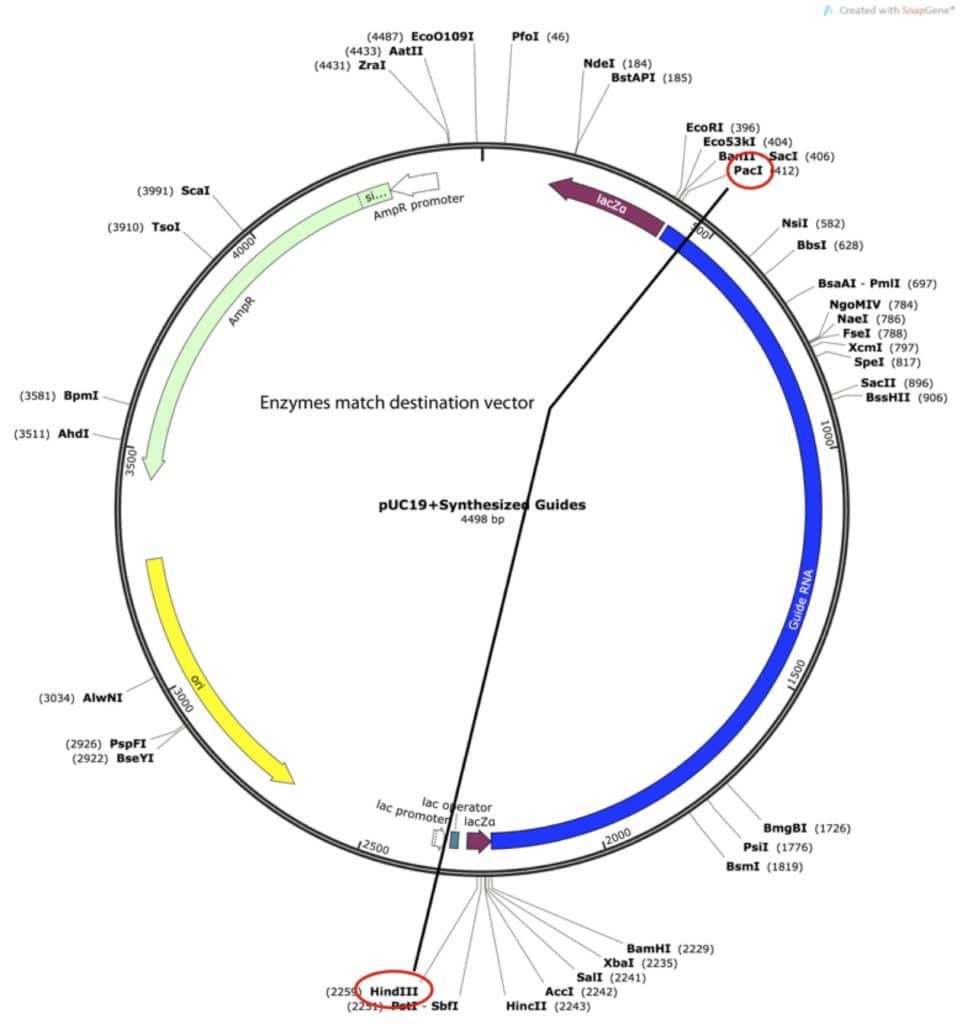
The guides are inserted into the MCS of pUC19, meaning that I can now cut them out with HindIII and PacI. Since HindIII and PacI are the enzymes already left behind in my Cas9 vector the cloning experiment is quite simple, it’s a simple cut and paste. So finally I am at the end product:
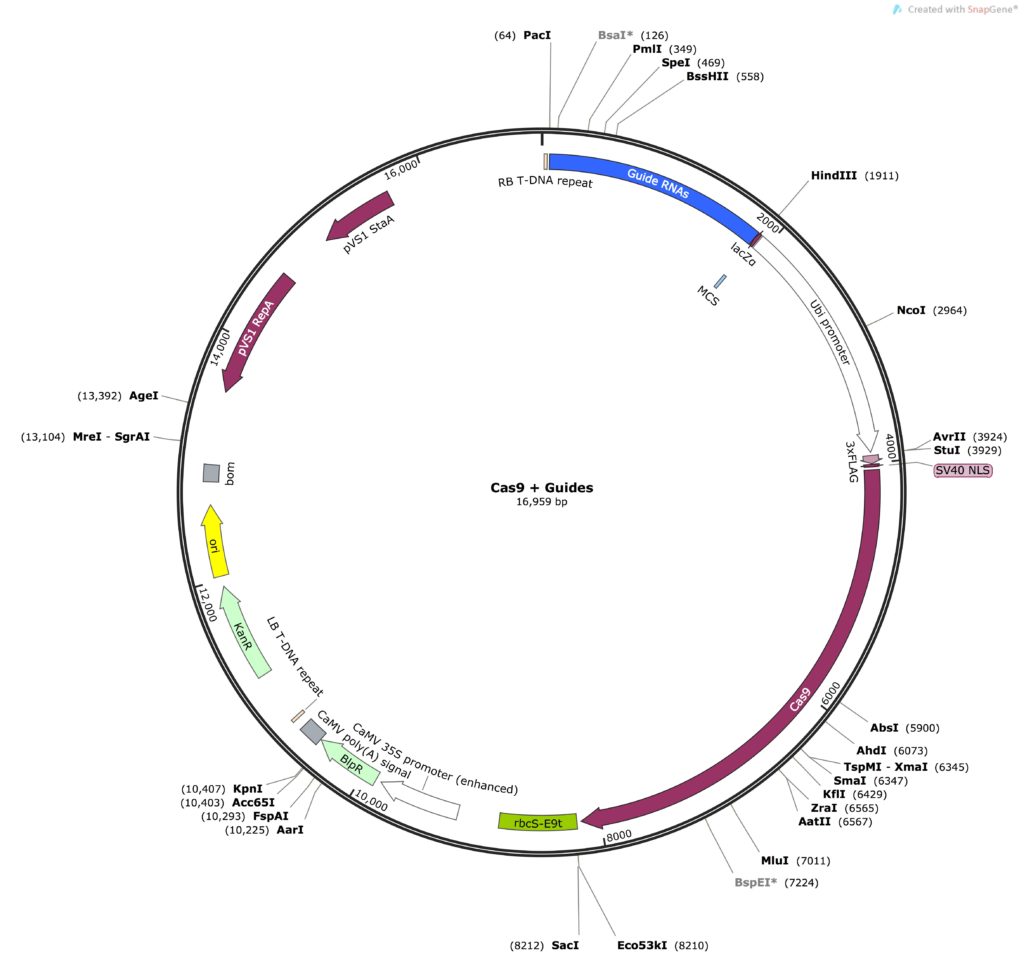
Tada!
Confirm it’s What You Want
Now that you have it all planned, make sure you met all of your original requirements. I know I keep reminding you of what seems truly obvious, but it comes from real experience. With the complexities involved in planning out a multi-step cloning experiment you really can lose focus of some of your minor requirements.
Document Your Restriction Cloning Plan
Now that you have your restriction cloning plan, document it. Save your files, and write down every component you need in order. SnapGene prints out a history summary, which I usually hang up over my lab bench to keep track of everything as I go. If your program doesn’t have something similar, I suggest making one of your own. Write down what enzymes you will use in each step, what the band sizes should be, the particular band you want to extract if doing a gel extraction, etc. I also like to make a list of primers I can use for verification while I have the sequence files up on the computer. This way I will have all of the information I need in one location.
The key to planning a restriction cloning experiment is to consider all of your options. Oftentimes you are restricted by available enzymes, or reliable primer binding sites, and you need to change your original plan. The key is to not get immediately discouraged and think through what other options you might have. Remember, there are still problems that might appear when you are at the wet bench, but for the most part, the planning is half of the battle. Good luck!
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
download
The Molecular Cloning Cheat Sheet
CHEAT SHEET
Nuclear Extraction Protocol