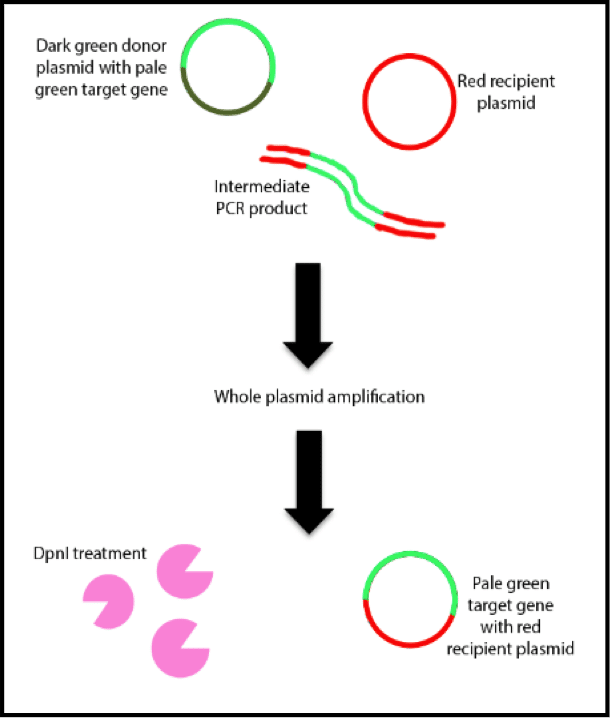
A while back I was doing a project where I had to make a mutation library of a plasmid. There are a number of ways to do this but I thought I would be smart. UV light is mutagenic – why not just put the plasmid on a transilluminator for a while? That should introduce some mutations and make my library with the minimum of fuss.
Well it was a reasonable idea, but the results were a real eye-opener.
I put aliquots of the plasmid into UVettes (plastic cuvettes that don’t absorb UV light) and exposed the aliquots to 302nm UV light for 0, 15, 30, 60, 120 and 180 seconds. I then transformed 1 microlitre of each aliquot into competent cells and plated on selective plates.
Put this article into practice
Choose a free resource to help you move forward
PROTOCOL
Chemically Competent Cells Protocol
CHEAT SHEET
Nuclear Extraction Protocol
I hoped to see a “killing” curve where the number of transformants I obtained would fall due to increasing damage to the antibiotic resistance cassette or the replication origin as the UV-exposure of the plasmid increased. Then I would pick an exposure time somewhere in the middle of the curve – one that caused enough but not too much damage – to do the real mutagenesis.
Well, I got the curve alright, but it turned out to be more like a flat-line.
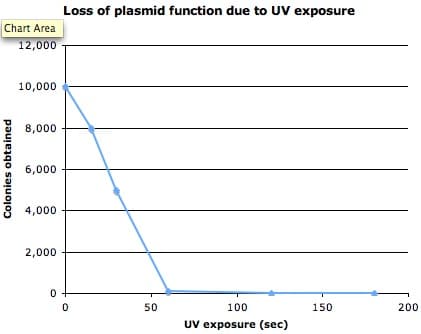
Ok, so this was at 302nm – a shorter and therefore more damaging wavelength than you would normally use for visualizing DNA you want to use later – but I found the level of damage sustained by the plasmid in such a short space of time startling.
Remember, the DNA fragments you use in cloning are in a sense more fragile than the plasmid since not only UV damage to the resistance cassette/origin of the plasmid, but also damage to the DNA ends, will scupper your experiment.
So, I’m sure you can see why this was an eye-opener. Like many people, I routinely used UV to visualize restriction digest bands that I then extracted and ligated in cloning experiments. And, like many people I wondered why it often didn’t work. But now the answer seemed clear – I knew that UV was damaging, but I didn’t realise it was that damaging!
I quickly switched away from using ethidium bromide for preparative gels and used methylene blue, a visible range strain, instead. This worked well – my cloning efficiencies increased immediately. This is a cheap solution, but it the procedure is quite time consuming since it requires long incubations in the stain.
If you’d prefer to cut down the staining times (warning: this means less excuses for coffee breaks), you could use stains such as SYBR-safe that can visualized with non-UV light. SYBR-safe is more expensive that EtBr and using it requires an initial outlay for a blue light box, but it is as sensitive and fast as fast EtBr, without the requirement for damaging UV. (Note: I don’t buy the whole EtBr scare story, but for purely practical reasons SYBR-safe is a good option in preparative gels.)
Or if you are intending to journey away from the (UV) light, and money is no object, you might want to do it in the Rolls Royce of gel systems – Invitrogen’s CloneWell. CloneWell gels are pre-cast (so even less excuses for coffee breaks) and have SYBR-safe incorporated in them. Using the E-gel holder’s built in blue light you can watch your DNA migrate in real-time, but the best thing about the CloneWell system is that the gels have a well halfway down the gel that you can use to collect your fragment without the need for gel extraction. So no UV and no gel extraction – sounds pretty good to me.
So, whichever method you use for DNA visualization, I would definitely recommend that you see the light and don’t use UV for preparative gels.
What do you use for DNA visualization?
Photo: james thrift
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
Nuclear Extraction Protocol
PROTOCOL
Chemically Competent Cells Protocol