Estimates indicate that there may be up to 2 billion living species of organisms, each with conserved and unique biological mechanisms that are vital for survival. How do scientists understand them all? Enter model organisms. Model organisms, as the name implies, are living things which are used as representative models for understanding other organisms. They are especially useful for investigating biological mechanisms. Thanks to model organisms, we now understand complex processes ranging from aging to zygote development.
Typically, model organisms are easy to grow, have short generation times, and are well characterized, with genes that are easily manipulated. Microorganisms are uniquely well suited as model organisms as most can be cultivated easily in the laboratory and have short generation times (minutes and hours compared to weeks and months for taxonomically higher organisms), making them the favourite subjects of in-depth investigation.
Rock-Star Microbial Model Microorganisms
In the A-list of model microorganisms, Escherichia coli and Saccharomyces cerevisiae are the rock-stars. E. coli, named after the microbiologist Theodore Eschrich who discovered it in 1884, is a Gram-negative bacterium commonly found in the gut of warm-blooded organisms. It is the most intensively studied free-living organism, with 12 Nobel prizes having been awarded to research that utilized this bacterial poster child. Indeed, the processes of transcription, translation, and replication, as well as the genetic code, were all first deciphered in E. coli, making the bacterium the most essential microorganism in molecular biology (Blount 2015).
Like E. coli, S. cerevisiae, also known as baker’s yeast, is frequently used in scientific research. S. cerevisiae was also the first fungus to have its genome sequenced. As many yeast proteins are similar in sequence and function to those in higher eukaryotes (Galagan et al 2015), including humans, studies performed in S. cerevisiae can help us to understand the functions of eukaryotic genes.
The Spotlight Shines on Emerging Model Microorganisms
Given the biodiversity of life, no one organism is a perfect representation of another. As we seek the answers to more questions in biology, other exciting but lesser known microorganisms have emerged as models. Examples of these emerging model microorganisms from each microbial classification are provided below.
Algae
Human diseases involving cilia (i.e. polycystic kidney disease) are often studied using algae such as Chlamydomonas because the flagella of algae share many similarities with cilia (Pazour et al, 2000). Algae have also been used to investigate mechanisms such as the permeability of membranes to water, osmoregulation, and salt tolerance.
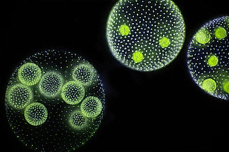
A striking example of a model alga is Volvox. This mesmerizing microalga forms spherical colonies of up to 50,000 cells, making it a common image on the covers of many textbooks. This beautiful ball of cells transitioned to multicellularity relatively recently in the tree of life (Russell et al, 2017) and is used as a model to investigate how single cells evolved to become multicellular organisms. Like other model organisms, Volvox can be transformed with exogenous DNA (e.g. transgenes). Knockdowns can also be performed to study the functions of genes. Due to their large size, Volvox colonies are easily seen under the microscope and mutants can be screened based on visual inspection.
Prokaryotes
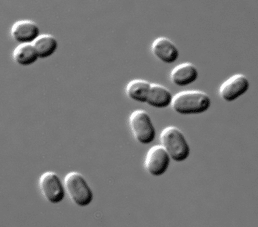
Bacteria such as E. coli were amongst the first microorganisms used as models due to their simple cell structure and ease of cultivation. Synechococcus is a cyanobacterium (photoautotrophic and can capture light through photosynthesis) that is found in marine environments. It has been used as a model organism to study a variety of cellular processes including circadian rhythms, carbon concentrating mechanisms, response to environmental stresses, photosynthesis, and cell division. The genomes and proteomes of some Synechococcus strains have been sequenced (Mackey et al, 2015; Coutinho et al, 2016), facilitating the understanding of functional and metabolic cellular processes.
Fungi
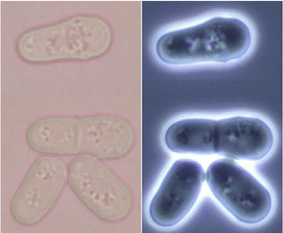
Like its budding yeast cousin S. cerevisiae, fission yeast – which reproduce by dividing themselves into two daughter cells of equal size – have come into the spotlight as a model microorganism. Fission yeast species Schizosaccharomyces pombe and Saccharomyces japonicus are used to study molecular processes such as responses to DNA damage and DNA replication. Many genes responsible for cell division and cellular organization in fission yeast are also found in the human genome, making it an excellent model for the investigation of these processes in humans. The genome of S. pombe has been fully sequenced, and sub-cellular localization of almost all the proteins in the yeast has been elucidated by tagging with Green Fluorescent Protein (GFP) (Matsuyama et al, 2006).
Protists
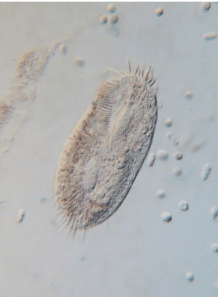
Being eukaryotes, many cellular features of protists and human cells are conserved, making them good representatives for studying essential processes that occur in human cells. Ciliates and amoebae are common model organisms from this classification. The curious trumpet-shaped giant ciliate Stentor coeruleus has a remarkable ability to regenerate itself from fragments, making it a model organism to study cellular regeneration and wound healing. The ciliate Oxytricha is used to investigate chromosome biology, post-zygotic development, epigenetics, and genome rearrangement. Naegleria gruberi exists both as an amoeba which crawls and a flagellate that swims. It is used to understand the cell motility of these two different states. Insights from these findings shed light on disease mechanisms such as the spread of cancer.
As we uncover more about the myriad of life and with the advancement of sequencing technology, it is likely that every organism can be a model. More intriguing microorganisms will likely emerge as models in the future. The possibilities of what we can understand from them are endless!
References
- Blount, ZD (2015). The Natural History of Model Organisms: The unexhausted potential of E. coli. eLife 2015;4:e05826.
- Coutinho F, Tschoeke DA, Thompson F and Thompson C (2016). Comparative genomics of Synechococcus and proposal of the new genus Parasynechococcus. PeerJ. 4: e1522.
- Galagan JE, Henn MR, Ma L-J, Cuomo CA and Birren B (2005). Genomics of the fungal kingdom: Insights into eukaryotic biology. Genome Res. 15: 1620-1631.
- Mackey KRM, Post AF, McIlvin MR, Cutter GA, John SG and Saito MA (2015). Divergent responses of Atlantic coastal and oceanic Synechococcus to iron limitation. PNAS. 112 (32): 9944-9949.
- Matsuyama A, Arai R, Yashiroda Y, Shirai A, Kamata A, Sekido S, Kobayashi Y, Hashimoto A, Hamamoto M, Hiraoka Y, Horinouchi S and Yoshida M (2006). ORFeome cloning and global analysis of protein localization in the fission yeast Schizosaccharomyces pombe. Nat. Biotechnol. 24: 841–847.
- Pazour GJ, Dickert BL, Vucica Y, Seeley ES, Rosenbaum JL, Witman GB and Cole DG (2000). Chlamydomonas IFT88 and its mouse homologue, polycystic kidney disease gene tg737, are required for assembly of cilia and flagella. J Cell Biol. 151(3): 709–718.
- Russell JJ, Theriot JA, Sood P, Marshall WF, Landweber LF, Fritz-Laylin L, Polka JK, Oliferenko S, Gerbich T, Gladfelter A, Umen J, Bezanilla M, Lancaster MA, He S, Gibson MC, Goldstein B, Tanaka EM, Hu C-K and Brunet A (2017). Non-model model organisms. BMC Biology. 15:55.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
PROTOCOL
Chemically Competent Cells Protocol
POSTER
Antibiotics Reference Guide