In situ zymography (ISZ) is the best choice for studying proteases. Proteases are a challenge to study as proteases are extremely potent enzymes. As such, they need to be controlled at multiple levels to prevent them from being unleashed and making a cellular mess. Regulation of their activity occurs at virtually all levels: transcriptional control, mRNA stability, post-translational modification, activation of latent proenzymes, spatial localization of secretion, and inhibition with endogenous tissue inhibitors.
So, herein lies the problem. Genetic techniques do not guarantee that translation takes place. Immunological techniques can detect an enzyme, but they usually don’t distinguish between the latent and the active form of an enzyme. Gel-zymography indicates enzyme activity, yet it is still powerless to tell whether the enzyme is blocked by its inhibitors in the cell. Immunohistochemistry can localize the inhibitor, but it can’t tell whether it is bound to the protease. Alas, after combining the three main approaches, we still don’t know where our protease does its chopping! But don’t despair! ISZ can save us.
The Principles of In Situ Zymography
Here’s how it works. Use a substrate specific for a protease in question that is modified so that upon digestion it is readily visible. Place the substrate on a tissue section. Then incubate the sample at an optimal temperature in a humidity chamber, which allows digestion of the substrate by the activated enzyme in its native location. View the lysis of the substrate by microscopy.
Sounds fairly simple, but if you want to get it right, you need to watch out for a few things:
Tissue Fixation
Neutral-buffered formalin is a general fixative for most purposes, but for ISZ, this is something to avoid. Formaldehyde forms crosslinks with functional groups of proteins, thereby inhibiting protease activity. There are two ways to overcome this problem. You can either use:
- Unfixed frozen tissue (cryosections)
This works generally fine, however, frozen sections often come with impaired tissue morphology, which makes microscopy difficult, and image quality poor. On the other hand, you do get to skip the time-consuming and smelly xylene bath and graded ethanol rehydratation steps, and can just air-dry the sections before further treatment. - Fixatives based on ethanol or zinc
Unlike formaldehyde, ethanol is not a crosslinking agent but fixes tissue by precipitation. Alternatively, zinc-based fixatives can also do the job. These fixatives leave the functional and morphological properties of the tissue intact, providing conditions for proteolytic activity as well as for good-quality imaging.
Visualization
There are a few different approaches that are useful to visualize the proteolytic reaction.
- Place a photographic emulsion containing the substrate over the slide containing the tissue section. Enzymatic activity using light microscopy appears as white spots on a dark background. This is an older technique, not in use much today due to limited sensitivity.
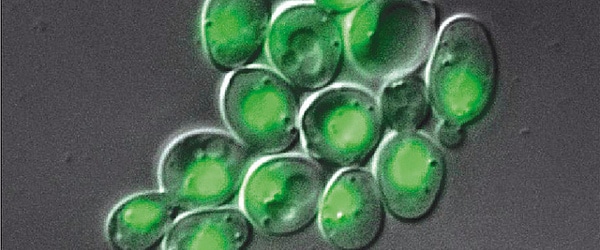
- Use a fluorescently labeled substrate and then detect enzymatic activity as dark spots against a fluorescent background using fluorescence microscopy.
- The substrate is dye-quenched (DQ). For example, DQ-gelatin improves the sensitivity in detecting matrix metalloproteinases-2 and -9. Fluorescein isothiocyanate (FITC) labels the gelatin so that the fluorescence is quenched. Cleavage by gelatinases produces fluorescent peptides, visible against a faintly fluorescent background.
Controls
Yey! We have a signal!
But, wait… How can we be sure that it comes from the exact protease we are looking for? – Our boring inner skeptic nags. Just to be on the safe side, we should probably introduce a few controls.
- Background noise
To estimate the background noise, add the substrate and then immediately place the tissue sections at -20°C for 2 hours. Temperatures that low block all enzymatic activity, ensuring you that whatever signal you see, surely does not come from a protease. Due to a tissue’s optical properties, you will probably get some signal on the control section as well, but it should be very weak.
- Fluorescence intensity
Set the signal-to-noise ratio on your microscope, all then examine all samples and controls under the same attenuation level. To ensure this, it’s a good idea to counterstain the nuclei with DAPI, and then merge the pictures from the green and the blue filter. The fluorescence from DAPI should be the same between the samples and the controls, while the fluorescence coming from FITC should vary. Counterstaining with DAPI also enables a better perspective on signal localization and tissue morphology.
- Contribution of other enzymes
But what if another protease has the same preference for the substrate as the one you are studying? If this is the case, it is essential to block all the other enzymes that can contribute to substrate degradation. Simply pre-incubate control slides with selective inhibitors.
And a Closing Thought
Once we have taken all these precautions, we have a quick, reliable, low-cost technique for detecting protease activity. A major drawback, though, is the difficulty in quantifying the results, and consequently, comparing samples. To get quantitative results use gel-zymography in parallel, and measure lythic zones by densitometry.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
POSTER
Immunofluorescence Troubleshooting Guide
POSTER
Histological Stains Poster