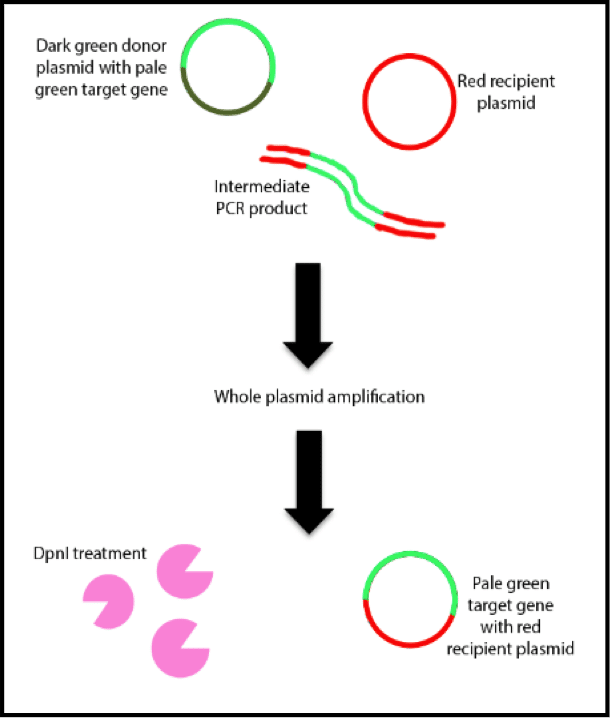
While the classic approach to molecular cloning – using restriction enzymes to excise a DNA fragment of interest – is as useful as ever, new techniques that make cloning faster, easier and more versatile are available.
As a smart molecular biologist, you should be examining each of them to see whether or not adding them to your tool-kit would be worthwhile.
One such technique is PCR cloning. This article is intended to help you decide if PCR cloning is for you.
What is PCR Cloning?
PCR cloning is a method that enables a PCR-amplifed insert to be cloned directly into a linear vector (either a ready-made commercial prep or home-made by PCR-amplification). After the ends are modified to make them compatible, the fragments are ligated and transformed as normal (for a more thorough explanation, check out this guide from the NEB website).
Put this article into practice
Choose a free resource to help you move forward
PROTOCOL
Chemically Competent Cells Protocol
CHEAT SHEET
Nuclear Extraction Protocol
No restriction enzymes = fewer purification steps
Traditional cloning of PCR products relies on restriction enzyme recognition sequences inserted in the primer sequence. This of course means that purification steps are required to remove the DNA polymerase before digestion to avoid further end modification or possible degradation following the restriction step. PCR cloning is a way to jump over these hurdles, as the PCR product can be directly used for cloning. This lack of purification steps makes the PCR cloning protocol very quick compared to traditional cloning, which is one of the main benefits of the technique.
Joining the ends
Once you have the amplified insert (and amplified/prepared vector) the next step is to join them by ligation. Depending on which DNA polymerase was used for the PCR, the insert’s ends may be blunt (if a proof-reading polymerase was used) or possess a single non-template dependent nucleotide (if Taq was used).
The nature of the ends determines the next processing steps for successfully cloning the amplicons, since the insert and vector must both possess complementary ends (i.e. both blunt, or both sticky). To match the insert ends and vector ends you can use specialized vectors for each kind of insert end, or carry out subsequent steps to convert both kinds of amplicon ends to match the ligation strategy of the vector being used.
Or alternatively you can use a kit that helps you get around the problem. A couple of examples are:
1. A ligase-independent approach to PCR amplicon cloning, in which ligation is carried out by a topoisomerase covalently bound to the (commercially prepared vector). Examples include StrataClone PCR kits from Agilent and TOPO® PCR cloning kits from Invitrogen™. These are powerful techniques. Although they come with the slight drawbacks that a separate kit is required for each kind of end, and insert ends must be unphosphorylated.
2. Another approach, from New England Biolabs®, uses a master mix containing prepared vector, invisible end-polishing components and ligase. These end-polishing components become active only when needed, if an insert has a single-base overhang (like those left by Taq), and quickly convert the ends to ligation-ready blunt ends. The master mix further streamlines the protocol, removes the need for specialized vectors for each type of end, and it can also be used with both phosphorylated and non-phosphorylated insert ends, with or without purification.
PCR cloning is non-directional
A further characteristic of PCR cloning is that inserts are ligated non-directionally into the vector. This can be an advantage or a disadvantage, depending on the next steps following cloning. But if the advantages of PCR cloning are attractive, you can of course overcome this problem with a restriction/sequencing screen for clones where the insert is in the desired direction.
But one application where non-directionality is attractive is if you are trying to clone an insert that is toxic for E. coli, as survival is possible for clones where the insert is introduced in a reverse orientation, and so is never translated.
Clonal selection with suicide vectors
Another time-saving approach inherent in some PCR cloning systems involves the use of “suicide vectors” to control colony development after transformation. These vectors include a toxic gene that, upon successful ligation, is interrupted by the insert, allowing the cell to survive. If the vector simply re-closes – which means unsuccessful ligation of the insert – the cell dies, and no colony is formed. This approach greatly simplifies downstream screening of the transformants as, with most suicide vector approaches, >95% of the colonies will have the insert.
What are the downsides?
Of course, PCR cloning is not a panacea. Like everything, it has its drawbacks and limitations. To get the maximum speed and versatility of PCR cloning, you’ll need to use a kit – which means a higher cost per experiment than traditional cloning, and also a more limited choice of vectors. If you need to clone multiple fragments in the same construct, PCR cloning is not for you either, as the lack of end-discrimination makes this problematic. And, as mentioned above, the non-directionality of the cloning may (or may not) rule it out for your application.
It’s all about saving time
But PCR cloning is a technique to consider if quick, high-efficiency cloning is your priority. The simple and versatile workflow allows for an extremely fast method – only a couple of hours from start to finish – ideal for projects with a high throughput, which would be difficult to achieve with more traditional methods.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
PROTOCOL
Chemically Competent Cells Protocol
CHEAT SHEET
Nuclear Extraction Protocol