ChIP-seq is a wonderful technique that allows us to interrogate the physical binding interactions between protein and DNA using next-generation sequencing. In this article, I’ll give a brief review of ChIP and introduce the chromatin immunoprecipitation sequencing technique (ChIP-seq), which combines ChIP with next-generation sequencing.
What is chromatin immunoprecipitation?
Chromatin immunoprecipitation (ChIP) allows us to determine protein-binding sites on DNA. Chromatin is the complex of DNA packaged with histone proteins into nucleosomes. ChIP makes use of reversible cross-links made between DNA and associated proteins by formaldehyde fixation of cells or tissue. The fixed chromatin is physically sheared and DNA fragments associated with a particular protein are selectively immunoprecipitated and analysed. Analysis can be on a locus-by-locus basis using PCR, but more commonly ChIP is interrogated with microarrays (ChIP-chip) or next-generation sequencing (ChIP-seq).
How does ChIP-chip work?
ChIP-on-chip, or ChIP-chip, combines chromatin immunoprecipitation with microarray (chip) analysis. In 2001, Jason Lieb in Pat Brown’s lab at Stanford published the first ChIP-chip paper (Lieb 2001). In this method, the DNA fragments that precipitate with a particular protein are applied to a microarray chip for analysis. This generates a global picture of where the protein binds, in contrast to simply interrogating single binding sites by PCR. While this was a revolutionary approach, the technique is limited by the array technologies used. Firstly, you need a microarray for the genome you want to study, and in the early to mid 2000’s this was often not the case. Secondly, those microarrays are limited by the quality of the reference genome and the ability to design probes that will work on an array. Thirdly, arrays are limited in how much of the genome can be represented, and usually tile across the genome at 100’s or 1000’s of bp resolution. There are also issues about bias in amplification of the ChIP DNA fragments, normalisation of array data, and comparability of array platforms.
How does ChIP-seq work?
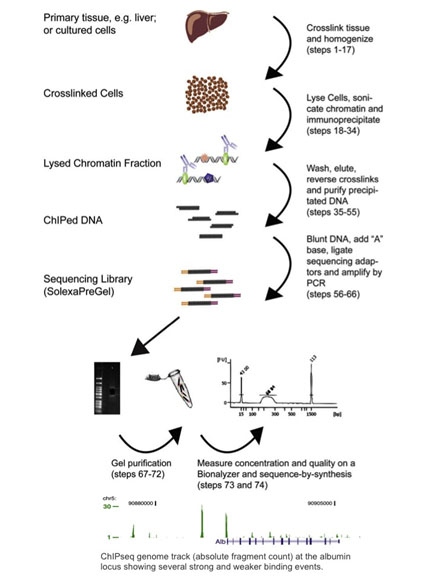
Chromatin immunoprecipitation sequencing, or ChIP-seq, combines ChIP with next-generation sequencing (Barski 2007, Johnson 2007). ChIP-seq protocols have been adapted from ChIP-chip methods: proteins are cross-linked to their bound DNA by formaldehyde treatment, cells are homogenized, and chromatin is sheared and immunoprecipitated with antibody-bound magnetic beads. The immunoprecipitated DNA is then used as the input for a next-generation sequencing library prep protocol, where it is sequenced and analysed for DNA binding sites. See the figure below for a summary of the ChIP-seq workflow, and an example of ChIP-seq results (reproduced with kind permission from Dominic Schmidt (Schmidt 2009)).
Enjoying this article? Get hard-won lab wisdom like this delivered to your inbox 3x a week.

Join over 65,000 fellow researchers saving time, reducing stress, and seeing their experiments succeed. Unsubscribe anytime.
Next issue goes out tomorrow; don’t miss it.
Although the majority of the 400 or so papers published so far have been analysed on the Illumina platform, ChIP-seq can be performed on any next-generation sequencer (Wold 2008). ChIP-seq has been widely adopted since it was first reported in 2007. In fact, it has almost totally supplanted ChIP-Chip, since it allows genome-wide analysis and does not have the limitations discussed above.
What can you actually do with ChIP-seq
ChIP-seq is a powerful tool and versatile tool, and there are many great examples of ChIP-seq use in the literature. I have picked a couple of my favourites (from work performed in the core facility I manage) to illustrate what is possible, and have included examples where ChIP-seq has inspired the development of new methods:
- Antoni Hurtado, et al. performed knock-down of the FoxA1 “pioneer factor”, resulting in reduced binding by the estrogen receptor (ER) at over 50% of known ER binding sites. They showed that FoxA1 is an important regulator of ER-mediated transcription, suggesting it may be a new and important therapeutic target in breast cancer (Hurtado 2011).
- Dominic Shmidt, et al. used ChIP-seq to investigate the evolution of transcription factor binding. They focused on CEBPA and HNF4 binding in the liver tissue of five vertebrate species: human, mouse, dog, opossum and chicken. ChIP-chip would have been almost impossible given the different species involved and complexities in designing probes (Schmidt 2010).
Further modifications to the ChIP-seq method have led to several new methods, including ChIP-seq for RNA-protein interaction analysis, as well as DNase-seq and FAIRE-seq, both of which are used to identify regulatory regions in DNA.
In summary, ChIP-seq is a mature technique that is preferable to ChIP-chip and can be used by almost any group with an interest in DNA:Protein interaction analysis. How could you use ChIP-seq in your research?
References:
Johnson et al, Genome-wide mapping of in vivo protein–DNA interactions. Science (2007).
Wold & Myers, Sequence census methods for functional genomics. Nat. Methods (2008).
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.