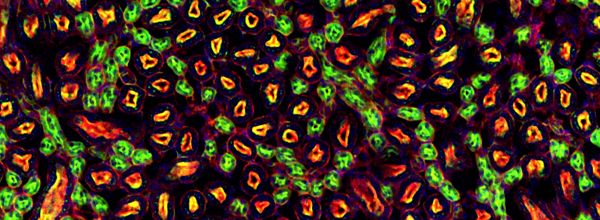
Are you trying to figure out how to calculate intensities of fluorescently-labeled single cells? Do you have cells at high densities or present in clusters? Are you worried that your current cell imaging analysis software is unable to mark clear boundaries around each cell in a cell cluster? Don’t fear, because CellProfiler 2.1 is here to help! From counting cells to quantifying fluorescence, it helps you distinguish cells from each other, especially if multiple cells are packed together in your microscopic image.
It’s an Easy-Peasy Tool for Cell Imaging!
CellProfiler 2.1 is a user-interactive, open source cell imaging analysis tool. There are various individual modules performing specific tasks, which can be put together to generate an image analysis pipeline. Users can then modify module parameters to generate the desired result. It is also really flexible, as it works on multiple platforms, such as Windows, Mac, and Linux. Furthermore, CellProfiler’s design is suitable for the high-content screening of thousands of images in batch modes on clusters.
What is the Fuss About Image Segmentation?
The performance evaluating step for any image analysis pipeline is dependent on the algorithm you chose for optimally segmenting an image. In particular, segmentation clearly demarcates the boundaries between objects or cells, by distinguishing between the foreground and the background. Lastly, this is an important task, especially when you are trying to quantify single-cell based image measurements.
Example: How to Count Labeled Bacterial Cells
Step 1
Mark the cell boundaries, using the IdentifyPrimaryObject module which detects the brightest objects (e.g. nuclei) against a dark background. It detects the darkest spots or centers of the cells, which are generally more uniform in staining, and is useful especially when cells are fluorescently labeled. To begin a new project in CellProfiler, select: File > New Project. Then to add a module, select: File > Add Module > Object Processing > IdentifyPrimaryObjects.

Figure 1: Adding IdentifyPrimaryObjects module to the pipeline.
Play around with the module parameters and optimize them to give you your desired results. The “?” button on the right gives you a detailed description of each particular parameter.

Figure 2: The list of parameters for IdentifyPrimaryObjects module.
Step 2
The IdentifySecondaryObjects module identifies and marks cell boundaries using objects it sees in the IdentifyPrimaryObjects module, aka Step1.

Figure 3: The list of parameters for IdentifySecondaryObjects module.

Figure 4: Output of IdentifySecondaryObjects module. Image on top is the original image and the image at the bottom shows the segmention image.
Step 3
The MeasureObjectIntensity module makes intensity measurements for each of the objects in previous modules. Then use the ExportToSpreadsheet module, to export the intensity measurements for each object to an excel file.
In Summary
CellProfiler provides a ‘test mode’ so you can compare several algorithms and choose the best module and settings to get the best result. Besides that, CellProfiler provides a range of accessible in-built data analysis tools, that link to external databases like Oracle or MySQL and external programs like MATLAB.
CellProfiler 2.1 makes your life easier so you don’t have to worry about your cells being at close together at high densities to take microscopic images. This makes your image quantifications easy and hassle-free. There is much more that CellProfiler does. So, now go play with it and explore!!
You can download CellProfiler 2.1 from: https://cellprofiler.org/.
Further Reading
Carpenter AE, Jones TR, Lamprecht MR, et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biology. 2006, 7(10):R100. doi:10.1186/gb-2006-7-10-r100.