If you have any training in genetics, chances are that during the course of your education you ran into those funny little sequences called microsatellites. These are repeated tandem motifs 1-6 nucleotides long, scattered all over our genomes. These used to be called “junk DNA,” because researchers thought that the repeats served no purpose. Nowadays, geneticists shy away from that term—studies have shown that these microsatellites are not just junk! They do serve a purpose, but there is still plenty of debate on what it is.
One thing everyone agrees on: microsatellites are highly polymorphic regarding their copy number. As such, they make pretty good genetic markers. There’s more good news! You can detect these small variations in allele length using chemicals and equipment that almost every lab has in stock. Stay with me, and I will show you how.
We Start with a PCR
First you need to amplify a region containing the tandem repeats. A standard 3-step PCR protocol should do the job. Design your primers so that they anneal to single copy DNA flanking the repetitive element. Several commercial companies that offer PCR kits specialized for microsatellite amplification. These can save you some time needed for optimizing the conditions. However, if you’re low on budget, you can do without them.
Running the Gel
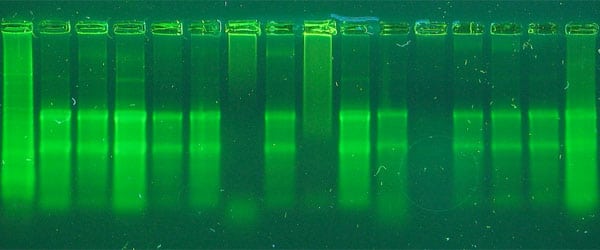
Most of us are use to resolving DNA fragments on native agarose gels. In this case, however, a denaturing polyacrylamide gel is a better choice. A 3%–20% polyacrylamide, depending on the size of the expected fragments, is optimal for separation of DNA less than 500 base pairs long.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
Nuclear Extraction Protocol
PROTOCOL
Chemically Competent Cells Protocol
In theory, you could run the gel in the absence of denaturants and keep the PCR product double-stranded. However, practice shows that the size of fragments can be determined much more reliably in denaturing conditions. Supercoiling of the helix can affect, to a certain extent, the electrophoretic mobility of native DNA. Also, some sequences can cause bending of the rod-like structure of DNA, which makes these fragments migrate more slowly than you would expect.
So take my word for it, separate the chains during electrophoresis. Dilute your samples in a loading dye containing 95% formalin. Then, heat the samples to 95°C and, afterwards, swiftly place the tubes on ice. The heat separates the strands allowing the formalin to work. Formalin then saturates the hydrogen bonds and thus helps prevents re-annealing.
To prevent secondary structure formation during migration, you need a denaturant in the gel. Using 6–8 M urea is the best choice for polyacrylamide gels. Temperature greatly enhances the denaturing effect of urea. So, ideally, you should carry out the electrophoresis in a 60°C incubator. Alternatively, you can adjust the applied power to ensure the gel reaches the correct temperature.
The voltage should be relatively low, around 5 V/cm, give or take. Use TBE buffer instead of TAE, because it has higher buffering capacity for prolonged runs.
It is also critical to load an appropriate amount of DNA. Load too little DNA, and you won’t detect anything. However, if you overload your lanes, then you may see smearing on your gel image. Acrylamide gels have a relatively high sample capacity—up to 10 µg can be loaded per lane.
Staining the Gel
Staining with silver is very sensitive method for detecting single stranded DNA. Reportedly, it is more sensitive than ethidium bromide staining. Nucleic acids reduce silver cations to insoluble silver grains that are then deposit in the gel.
You can visualize the bands by soaking the gel in a solution of silver ions and a reducing agent. The bands gradually develop as dark brown regions against a fainter, but also darkening background. You can stop the reduction, when the contrast is at its the strongest, by altering the pH of the gel.
Estimating the Size
After staining the gel, we estimate the number of tandem repeats by comparing the bands with a low molecular weight DNA ladder. The ladder should go through the same denaturation procedure as the samples. Note: many ladders are composed of direct repeats. This can result in arcs or smears because of partial renaturation. If possible, you should use DNA markers prepared by restriction digests, which perform better.
This method is highly sensitive. It is capable of resolving fragments of 2 to 500 nucleotides with length differences as small as a single nucleotide.1 To increase precision in interpreting results, it is a good idea to also load previously sequenced samples with the unknowns, and compare the bands.
For some purposes, instead of calculating the absolute number of tandem repeats, it is enough for the authors to classify them as “low copy number” and “high copy number”. In this case, you can load a DNA fragment you define as a “cut off” standard.
So, if you are doing your research in conservational genetics, behavioral genetics, forensics or population structure analysis, this method may come in handy. Happy counting everybody!
References:
- Summer H, Grämer R, Dröge P. (2009). Denaturing urea polyacrylamide gel electrophoresis (Urea PAGE). J Vis Exp. 32: 1485. doi: 10.3791/1485
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
Nuclear Extraction Protocol
PROTOCOL
Chemically Competent Cells Protocol