Ever since the invention of the first compound microscope by Zacharias Jansen in 1590, our understanding of the microscopic world has grown exponentially. Microscopes have evolved from mere assemblies of magnifying lenses to extremely powerful tools for visualization on the atomic scale. You can find a wealth of information on the workings of a microscope and valuable tips to produce some great images in our microscopy and imaging archives. In this article, I will share tips for both image acquisition and processing and why getting them right is critical for publishing scientifically acceptable data.
Scientific and Photographic Image Processing
Throughout this article, I will be using the terms, ‘scientific image’ and ‘conventional photograph’ to illustrate the difference between a microscopy image and a photograph from a digital camera. There are a few aspects that one should keep in mind when working with scientific images. The goal of acquiring or processing scientific images is to ensure that the image addresses the hypothesis or query in question but not necessarily make artistic impressions. Sure, there are great examples of microscopic images that are truly masterpieces (here’s one example), but the intended purpose of scientific imaging is to document changes happening in a biological system as accurately as possible.
In a conventional digital image from a camera, one often has to perform a little post-processing to make the image stand out. Apart from the usual brightness, color and contrast settings, you might also want to apply some artistic effects in Photoshop or alter the color landscape of the image to make it more appealing. These changes alter the underlying pixel values, and this is not acceptable in a scientific image.
A scientific image on the other hand, requires post-processing of a different kind. Post-processing of microscopy images goes beyond simply changing colors or adding filters. It requires knowledge of image processing algorithms and the effects these will have on data integrity. For example, I can make the sky look bluer in a photograph but I cannot simply make a fluorescence channel look more saturated that I actually can, especially in gene expression studies.
Put this article into practice
Choose a free resource to help you move forward
POSTER
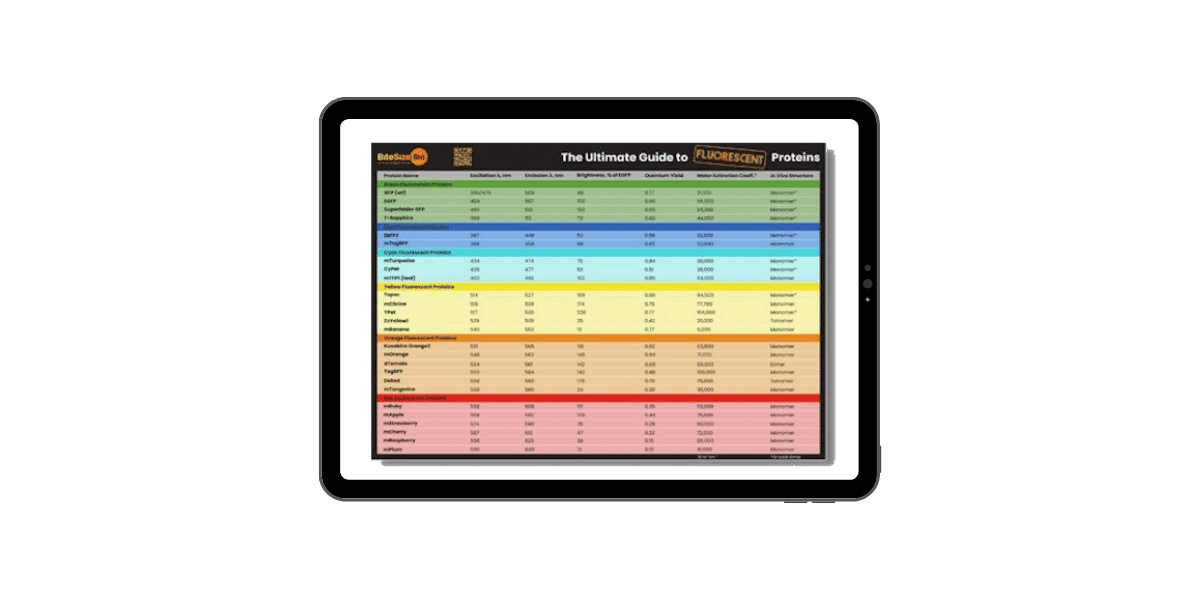
Fluorescent Proteins Guide
POSTER
Immunofluorescence Troubleshooting Guide
Cameras used in scientific imaging are far more sophisticated than conventional photographic cameras. Firstly, they are often cooled to very low temperatures to minimize electronic noise. Secondly, they support increase of pixel size by a process called binning. Furthermore, they are generally low on the megapixel count but feature larger pixels to capture the maximum number of photons possible. Cameras that are used for fluorescence-based imaging are usually monochromic as they devote all the available pixels to the all photons emitted from the sample instead of just individual red, green and blue pixels.
Acquire the Best Image, Every Time
Ideally, the need for post-processing of microscopy images should be as minimal as possible. This helps ensure that all relevant parameters are set up appropriately during image acquisition itself. For example, you might find the need to reduce noise during post-processing. While it is possible to deconvolute the original image and still get an acceptable result, reducing noise during acquisition will reduce the number of processing steps. Increasing the intensity of the source lamp, increasing the laser power, or altering photomultiplier tube (PMT) sensitivity are some ways in which you can boost signal and reduce noise. Similarly, it is important to set the right saturation levels during acquisition. The saturation should be set based on the original monochrome image to ensure correct pixel intensities, and it should be kept constant for the remainder of the imaging experiment.
You can also minimize the need for background correction during acquisition itself by adjusting the camera shutter time or PMT voltage settings accordingly (this depends a lot on the sample). It is also important to always acquire at the full bit depth supported by the camera/microscope. An 8-bit camera can acquire 28 values of intensities (0-255), which is the common bit depth of conventional displays. Confocal microscopes can acquire upto 212 (0-4095) intensities or higher. The higher the bit depth, the higher the range of intensities between the whitest white and the darkest black – which translates to more accurate pixel representation. If you reduce the bit depth during acquisition by artificially changing the Look Up Table (LUT) values, you will have of loss of pixel information during analysis.
Avoid increasing camera/PMT gain unnecessarily. Increasing the gain increases both signal and noise together. Instead, try increasing camera exposure times or PMT voltage/laser power to increase signal and reduce noise. Also, be wary of increasing the offset voltage (negative current) beyond a certain threshold. There is a high chance that information from the sample gets lost if the offset value is too high – and no amount of post-processing can get the lost information back.
Ideally, avoid both zooming and magnification unless warranted as it can have a significant impact on image quantitation. If you need a magnified image, switch to a higher mag objective instead of applying software zoom or magnification.
You should also be considering minimal variability between experiments by ensuring more or less equal and at the same time, adequate warm-up times of the light source. Never switch setups between experiments and ensure that the microscope can detect the fluorochromes that you intend to use. If in doubt, discuss with your lab’s facility manager.
Finally, always save the originally acquired image in the microscope’s default raw image format. Raw image formats contain all the information captured by the camera sensor or PMT and also store valuable metadata pertaining to the acquisition. This helps you to quickly review or apply the same settings to later acquisitions, thereby maintaining consistency. The raw files also allow you to non-destructively edit the image. Often, you might need to export these raw files into common formats such as TIFF or PNG. Avoid lossy compression formats such as JPEG and always save images as TIFF files or if needed, PNG. TIFF is a lossless format that can also contain metadata and is read by most image viewing software.
Important Post-Acquisition Procedures
Even with the best acquisition practises, you will still find the need to do some post-processing. In this section, I will discuss some of the common post-processing procedures and the likely impact on your data. Like many microscopists, I recommend you familiarize yourself with the open-source software, ImageJ, or its fully loaded counterpart, Fiji. For an introduction to ImageJ/Fiji, and to image processing in general, you can refer to this article by Peter Bankhead.
Image comparison. More often than not, you might need to compare images across various timepoints e.g., during drug effect experiments. One of the easiest ways to visualize this information is to compare image histograms. Histograms depict the pixel information in terms of intensities or even as statistics. Even better is to identify specific regions of interest (ROI) such as the nuclear or ER compartment and compare the histograms across the ROIs of multiple images. But remember that histograms are not always the ideal comparison tools and non-identical images can still have identical histograms.
Multi-dimensional data. Multi-dimensional data is not just restricted to x, y and z planes, but can also include color and time information. An important aspect of post-processing is being able to exactly isolate the required information from the multi-dimensional dataset. You can also do maximum and minimum z-stack projections. Avoid comparing pixel intensities from z-projections. They are not the actual representation of the sample. Instead, you might want to isolate a specific stack as a 2D image and then go about calculating the intensity.
Multi-channel data. Multi-channel images contain red, blue, and green colors in each channel which are then merged to generate a composite image. One of the post-processing steps involved is to edit the channel information or study the merged image for co-localization of proteins in a cellular compartment. Co-localization can be confirmed by using Pearson’s correlation coefficient.
Background subtraction. As discussed above, it is always better to minimize the background during acquisition itself. However, it is not always possible to do so. During post-processing, you can remove the background information from the image or filter out noise. Be careful to set the threshold such that pixel information from the actual sample is not affected.
Measurements. Measurements are one of the prime reasons for image post-processing. You can measure a host of parameters in an image, starting from the intensity value of each pixel to advanced statistics and even FRET, FRAP, and diffusion measurements. Based on the measurements, you can also do image segmentation and thresholding to exactly quantitate the parameter you desire.
Annotation. Annotating images helps you to deliver more information to the viewer. For example, you can draw a scale bar to indicate relative sizes or overlay information about the cell or fluorochrome being investigated. While annotating, especially while overlaying scales, make sure that the image processing program is calibrated for the very specific objective that you have actually used to acquire the image. Remember that no two objectives, even from the same manufacturer, are the same.
Conclusion
Although both scientific images and conventional digital photographs are made of pixels, the way in which we go about utilizing the information is very different. As you can see, it is important to acquire the image keeping in mind the likely downstream post-processing steps. This ensures that you will spend minimal time enhancing the image and more time deducing scientific information.
Ultimately, understanding your microscope setup and the software that goes along with it is of prime importance in getting the most comprehensive scientific information out of your images.
Further Reading
Paddock SW. (1999) Confocal Laser Scanning Microscopy. Biotechniques 27(5): 992-6, 998-1002, 1004.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
EBOOK
Guide to Special Stains for Histology
POSTER
Immunofluorescence Troubleshooting Guide