PIPE PCR is a ligase-independent, restriction enzyme-free cloning strategy like SLIC (link to my SLIC article), SLiCE and CPEC. The PIPE method eliminates sequence constraints and reduces cloning and site mutagenesis to a single PCR step followed by product treatment. It is fast, cost-effective and highly efficient. The key step is designing the primers; one of our readers also recommended this very helpful tool to get you started.
What is PIPE PCR?
The Polymerase Incomplete Extension method may be used for cloning and mutatgenesis experiments. It is an effective method of making initial clones, mutant sequences and truncated genes and was originally designed to microscreen for constructs with high crystallization potential.
How does PIPE work?
Normal PCRs generate mixtures of incomplete extension products and it’s these guys that make PIPE PCR possible. By following certain primers design rules, the PCR process introduces short, overlapping sequences at the ends of these incomplete extension mixtures. This allows complementary strands to anneal and produce hybrid vector-insert combinations. These can be transformed directly into recipient cells without the requirement of post-PCR enzymatic manipulations. This in turn enables thousands of genes to be cloned in parallel. The method is flexible but robust and can be automated thanks to its simplicity.
There are three distinct components of the PIPE method. The I-PIPE, or insert PIPE, step is common to the PCR you’re used to. This step is where the inserted genetic material, or simply the insert, is amplified to create a single-stranded product. V-PIPE, or vector-PIPE, is the next step but this can be done congruently in a different reaction. This point is where the vector is cloned. These two steps are combined for the final step of PCR amplification to add insert sequences into the vectors. To add mutations in the vector, a third step may be added, M-PIPE, or mutagenic PIPE, which allows you to make inserts, deletions and variations of a sequence. In M-PIPE, the insert-vector hybrid is amplified in the presence of special primers, one of which is complementary to the insert and the other to the vector.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
Lab Math Cheat Sheet
DIGITAL TOOL
Lab Math Calculator
The Special Primers
So what do we want in PIPE primers? Firstly, PIPE PCR requires that the first 15 bases on the 5’ end of the primers must be designed to be directionally complementary such that the resultant PCR fragments can anneal as desired and become viable plasmids following transformation. Quite a mouthful! Let’s look at this more closely with a visual aid (Figure 1)
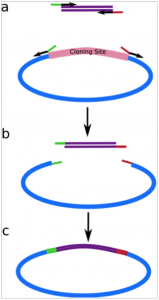
Figure 1. The PIPE cloning method. The insert and vector are amplified in separate reactions (I-PIPE (a) and V-PIPE (b)) and then mixed together to create vector-insert hybrids (c). Special primers can then be used to create mutations (M-PCR).
In I-PIPE PCR, the primers contain 5’ sequence complementary to the two distinct ends of the amplified vector and so the annealing occurs directionally to create you plasmid. In M-PIPE PCR the whole plasmid is amplified. The forward and reverse primers used must be designed such that a mutation is present and that they are complementary to each other. This creates a linearized PCR product that can self-anneal to form, viable, mutant plasmids.
The Steps Involved
Each Method Follows the Same Protocol
PCR is used amplify cloning vectors and inserts or for generating site-directed mutants. You’ll need to purchase PIPE Pfu mastermix. The PCR cycle design, the primers and the template DNA you use will depend on what you’re trying to do.
For V-PIPE: a recipient expression plasmid should be used, such as pSpeedET.
For I-PIPE: genomic DNA, PCR product or mini-prepped DNA from a clone you generated before or cDNA can be used for the insert. You can use any vector that can be amplified by PCR.
For M-PIPE: miniprepped DNA must be used unless you are certain that normal PCR will work for your construct.
To confirm your PCR worked, use gel electrophoresis to visualise the products.
Et voilà! Your plasmids are home free and ready to go!
References:
Klock HE and Lesley SA. (2009). The Polymerase Incomplete Primer Extension (PIPE) method applied to high-throughput cloning and site-directed mutagenesis. Methods Mol Biol. 2009;498:91–103.
Klock HE et al. (2008) Combining the polymerase incomplete primer extension method for cloning and mutagenesis with microscreening to accelerate structural genomics efforts. Proteins. May 1;71:982–94.
Kumar A et al. (2012) Structure of a novel winged-helix like domain from human NFRKB protein. PLoS One. 2012;7:e43761
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
Lab Math Cheat Sheet
DIGITAL TOOL
Lab Math Calculator