The ‘diffraction limit’ of a microscope is the minimum distance between two fluorophores where they can be still be discriminated as two separate objects. This diffraction limit has long constrained attempts by biologists to observe the intracellular environment. With a lower limit of ~200nm in confocal microscopy, this diffraction limit significantly limited the detail you could obtain when imaging cellular structures, and protein interactions and dynamics. But not anymore! This lower limit was shattered with the recent advent of some new techniques. So sit back and relax, as I tell you about the latest advances in optical resolution and open up to you a whole new world of fluorescence microscopy.
Last year, Justin wrote a great guide to super-resolution microscopy where he outlined the different techniques with their pros and cons. In this article (and a few more to come) I will go into more detail about these techniques, explain their importance and give some examples of how they are being used:
Total Internal Reflection Fluorescence (TIRF) microscopy
Before I talk about super-resolution microscopy techniques, however, I would be remiss not to mention TIRF. TIRF microscopy (also known as evanescent wave microscopy) is based on classical widefield microscopy, where fluorophores near the coverslip (and therefore near the cell surface) are selectively excited whilst fluorescence further away from the coverslip is minimised.
The technical stuff:
Put this article into practice
Choose a free resource to help you move forward
POSTER
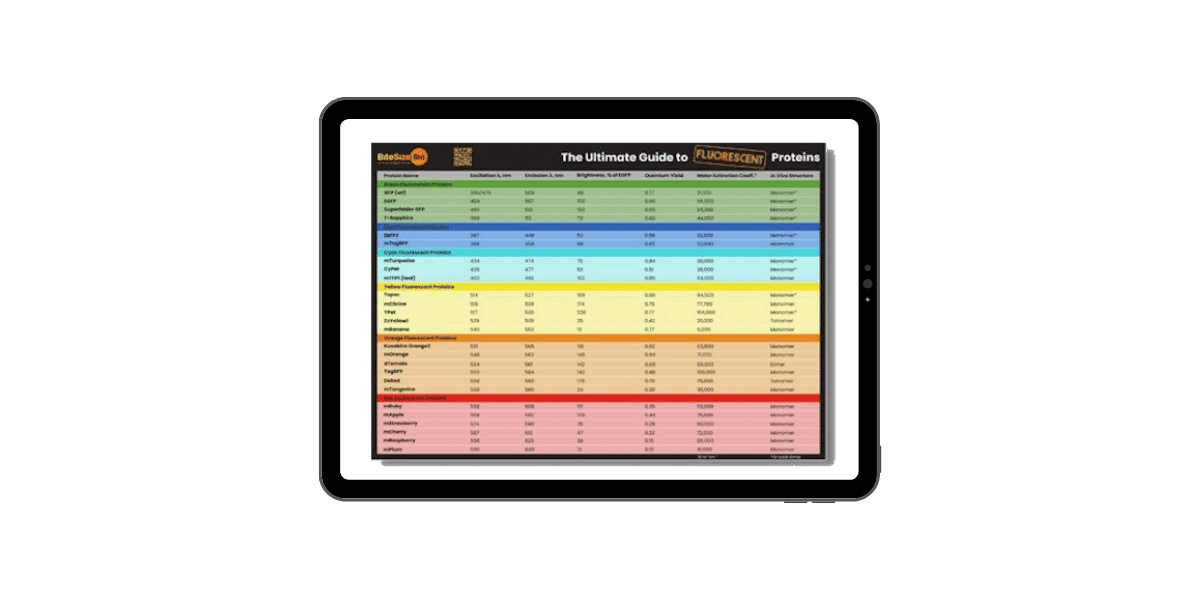
Fluorescent Proteins Guide
POSTER
Immunofluorescence Troubleshooting Guide
TIRF microscopy is achieved by taking advantage of the so-called critical angle of light, resulting in the light being totally reflected at the coverslip-sample interface. This results in the creation of an evanescent wave (a near field wave which decays exponentially) that can only penetrate ~100nm into the sample. Because only fluorophores within this wave are excited there is less background fluorescence, which makes TIRF sectioning depth superior to confocal microscopy
How does this help you?
TIRF microscopy works with both fixed and live cell samples. It can help you image membrane-proximal events at an improved resolution, whilst removing interference from fluorescence within the cell. In this way it can be used to image the cytoskeleton, membrane bound proteins and intracellular molecules which are close to the plasma membrane.
Examples of how it’s been used:
- Microtubules are responsible for golgi vesicle transportation (Schmoranzer and Simon, 2003)
- Imaging dynamics of signaling molecules at the T cell immune synapse (Purbhoo et al, 2010)
Super-resolution imaging techniques
Now let us enter the realms of ‘super-resolution’. These techniques can be, broadly speaking, grouped into two main classes:
Class 1: Ensemble techniques – where the improvement in resolution comes from minimizing the size of the point spread function (PSF; the smallest representative of a single object that the system can form) during image acquisition. Two main ensemble techniques are Structured Illumination Microscopy (SIM) and Stimulated Emission Depletion (STED) microscopy, which have very different approaches to reducing the PSF during acquisition (and therefore improving resolution)
Class 2: Single particle localisation techniques – where resolution is improved by localising single molecules.
Which technique is best for you depends on a combination of factors, including what you want to image and how you want to image it (live/fixed, etc). So first let us take a look at Ensemble techniques. Later this month I will cover single particle localisation techniques.
Ensemble technique #1: SIMS Microscopy
In June last year, Justin wrote a good explanation of how SIM works (including the maths!), but here is a brief overview to refresh your memory.
The technical stuff:
In essence, SIM uses patterned lines of light that are changed and rotated to create a series of images with ‘high spatial frequency’ information. By imaging the sample with structured light patterns in different orientations, the images can be combined and reconstructed into a high resolution image using computer algorithms
How does this help you?
- Theoretically, you can achieve a two-fold improvement in lateral resolution using this approach.
- SIM has become an attractive candidate for biological applications because standard dyes and staining protocols can be used, as long as the fluorophores used are stable enough to withstand multiple rounds of imaging.
Examples of how it has been used:
- Microtubule polymerisation and depolymerisation dynamics in mitotic cells (Kner et al, 2010)
- The mesh of actin filaments at the Natural Killer cell immune synapse is remodelled to allow lytic granules to pass through (Brown et al, 2011)
Ensemble technique #2: STED Microscopy
STED microscopy, on the other hand, uses a different approach to reduce the PSF, by de-exciting fluorophores that have already been excited. I will write a future article discussing STED microscopy in more detail, but for now here is a quick overview:
The technical stuff:
In this technique, the sample is illuminated with a laser at a focal spot, in the same way as in confocal microscopy, but this focal spot is surrounded by a doughnut-shaped beam formed by a laser of a longer wavelength (called the STED beam). This depletes the fluorophores surrounding the focal spot back to the ground state, resulting in a smaller spot being detected.
How does this help you?
- A resolution of 30-80nm can theoretically be achieved using this technique (Hell and Wichmann 1994).
- The technique is not axially limited, therefore cellular structures deep within the cell can be imaged. Live-cell STED imaging has also been reported.
It is important to note, however, that the choice of dye is crucial for STED microscopy, as the wavelength of the depletion laser should not overlap with the excitation range of the dye
Examples of how it had been used:
- Distribution of proteins at neuronal synapses (Willig et al, 2006)
- Organisation of SNARE proteins in the plasma membrane (Sieber et al 2006)
So that’s it for now on the ensemble techniques. In part 2 of this article, I will be discussing single molecule localisation techniques and how they differ from these ensemble techniques.
References:
Brown, A., Oddos, S., Dobbie, I., Alakoskela, J-M., Parton, R., Eissmann, P., Neil, M., Dunsby, C., French, P., Davis, I. and Davis, D. (2011) ‘Remodelling of cortical actin where lytic granules dock at natural killer cell immune synapses revealed by super-resolution microscopy.’ PLoS Biol.9 (9).
Hell, S. and Wichmann, J. (1994) ‘Breaking the diffraction resolution limit by stimulated emission: stimulated emission depletion fluorescence microscopy’. Optics letters 19 (11): 780-782
Kner, P., Chhun, B., Griffis, E., Winoto, L. and Gustafsson, M. (2009) ‘Super-resolution video microscopy of live cells by structured illumination.’ Nature Methods 6 (5): 339-342
Purbhoo, M., Liu, H., Oddos, S., Owen, D., Neil, M., Pageon, S., French, P., Rudd, C. and Davis, D. (2010) ‘Dynamics of subsynaptic vesicles and surface microclusters and the immunological synapse.’ Sci. Signal. 3 (121).
Schmoranzer, J. and Simon, S. (2003) ‘Role of microtubules in fusion of post-golgi vesicles to the plasma membrane’. Mol Biol Cell 14 (4): 1558-1569
Sieber, J., Willig, K., Heintzmann, R., Hell, S. and Lang, T. (2006) ‘The SNARE motif is essential for the formation of syntaxin clusters in the plasma membrane.’ Biophysical journal 90L 2843-2851
Willig, K., Rizzioli, S., Westphal, V., Jahn, R. and Hell, S. (2006) ‘STED microscopy reveals that synaptotagmin remains clustered after synaptic vesicle exocytosis.’ Nature 440: 935-939
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
POSTER
Fluorescent Proteins Guide
POSTER
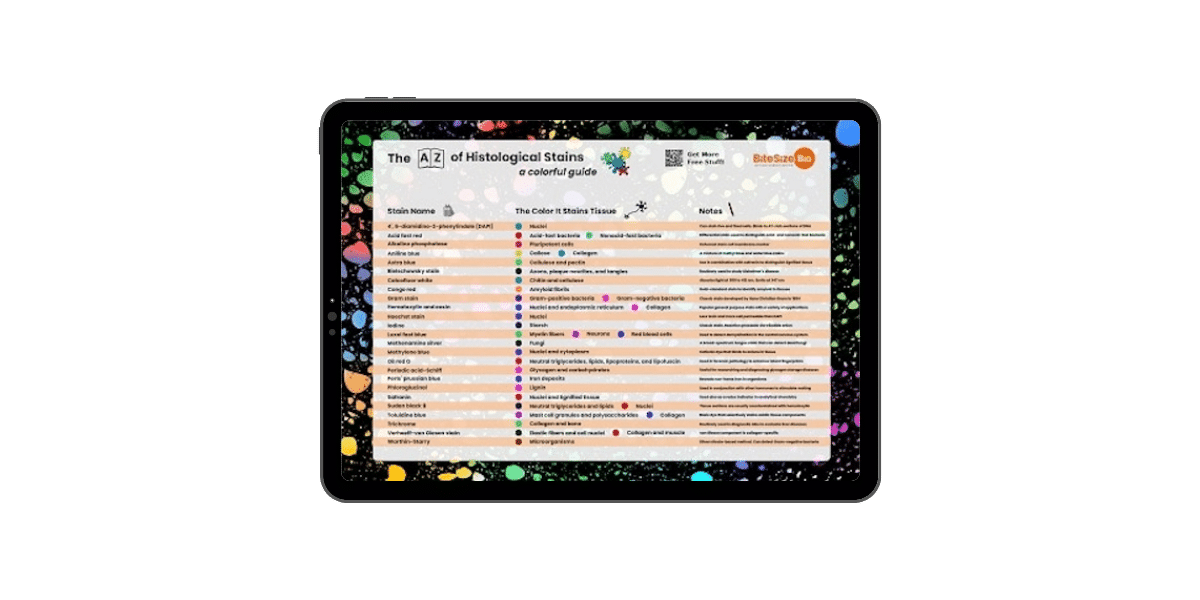
Histological Stains Poster