So you’re designing a new experiment that requires PCR quantification. You used to have only one method to choose from, but now you have two – Quantitative Real-Time PCR (qPCR) and Digital PCR (dPCR). Which one is right for your application? Both methods have good quantification, sensitivity and specificity for most applications. They are compatible with the same sample prep methods and use the same reaction components. How do you choose? Well as usual, the answer is in the details.
How are they the same? How do they differ?
Similarities
- Compatible with the same sample prep methods
- Amplification reaction components and concentrations are the same – PCR Master Mix, fluorescent probes, primers, nucleic acid target
- Both quantify the amount of target present in the sample
- Similar initial sample volumes
- Both work with either hydrolysis probe or DNA-binding dye detection
- Wide dynamic range
- Multiplexing capability
Differences
|
qPCR |
dPCR |
|
Requires a standard curve |
No standard curve required |
|
No sample partitioning |
Sample is partitioned into many wells |
|
Data is collected in real-time |
End point data collection |
|
Relative quantification |
Absolute quantification |
When is Quantitative PCR the best choice?
For many applications, qPCR is still the best choice and remains the “gold-standard” method for nucleic acid target quantification. If a larger dynamic range, cost or monitoring real-time reaction efficiency are considerations for your application, then qPCR is preferable to dPCR. Quantitative PCR is also the application of choice for relative gene expression if the differences are > 2-fold. Quantitative PCR currently has a larger menu of general applications. Current automation and throughput capabilities of qPCR are superior to dPCR, but that gap is quickly closing.
When is Digital PCR the best choice?
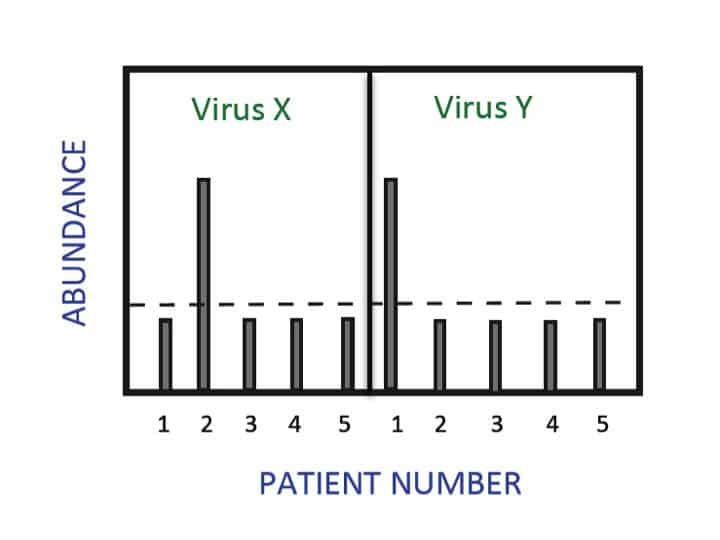
Although qPCR can be used for a wide variety of applications, there are certain aspects of those applications that make dPCR a better choice. Due to its use of sample partitioning and absolute quantification (instead of standard curve relative quantification), dPCR is best suited to applications requiring precision, high sensitivity and reproducibility.
Examples of these applications are:
Enjoying this article? Get hard-won lab wisdom like this delivered to your inbox 3x a week.

Join over 65,000 fellow researchers saving time, reducing stress, and seeing their experiments succeed. Unsubscribe anytime.
Next issue goes out tomorrow; don’t miss it.
- Rare allele detection
- Copy number variation
- Gene expression for <2-fold differences
- Quantification of NGS libraries
- Detecting low-abundance RNA
- Pathogen detection
- Viral load detection
The Generalist or the Specialist?
When do you choose dPCR vs qPCR for your application? Quantitative PCR has a wide variety of PCR applications and commercially available equipment, but it’s also been available for decades. Digital PCR is a relatively new technology with the first commercially available dPCR system introduced in 2006 by Fluidigm Corporation. There are many applications that are amenable to both dPCR and qPCR, with the number expected to grow for dPCR as the technology progresses. To differentiate their current applications, I like to think of qPCR as “the Generalist” and dPCR as “the Specialist”. While qPCR works well for many applications, there are certain subsets of those applications that require the added precision, sensitivity and reproducibility of dPCR.
Which method does your laboratory use and why?
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.