Most differential gene expression (DGE) experiments start with a simple question: How does expression change between different conditions?
Whether you are studying differentiation and development, or how cells respond to a potential drug candidate, answering that question is often constrained by practical logistics like budget and time. For many studies, this means choosing between the fast and accessible qPCR and slower, but more comprehensive methods like RNA-seq.
But now, the compromise between speed and scale is gone, with a new 3’ end counting RNA-seq service that makes it practical to gather transcriptome-wide data earlier in your experiments.
In this article, we will discuss the pros and cons of choosing qPCR or RNA-seq for DGE studies, and how 3’ end counting could forever change the way that you analyze gene expression.
The Distinguished, But Humble qPCR
Quantitative PCR has been the reigning technique for gene expression studies for decades, and for good reason. It is simple and effective, being a variation on the most essential tool in molecular biology: the PCR.
The main advantage of qPCR is its accessibility. Most labs doing gene expression studies own a qPCR machine, and the reactions need little more than standard PCR reagents (primers, polymerase, dNTPs, etc.) and a fluorescent probe for visualization.
Of course, qPCR shares the same technical challenges as any regular PCR, with some additional considerations. Since the objective is accurate fluorescence-based quantification, it is essential to limit off-target amplicons that would lead to erroneous measurements. This means that primers must be carefully designed, and reactions optimized, before reliable qPCR data can be collected.
The other, and ultimately more consequential drawback is that the qPCR workflow can only measure a handful of genes at a time, meaning that your insights are limited by pre-existing knowledge of the system, and qPCR experiments serve only to validate hypotheses rather than generate new ones.
Why RNA-seq Has Not Been Routine
A key advantage of RNA-seq, when you are deciding between qPCR or RNA-seq, is its ability to measure every mRNA transcript in a sample during a single NGS run.
However, traditional RNA-seq workflows can take 2-4 weeks or more for turnaround because of logistical bottlenecks. Library preparation, sample batching requirements, queue delays, and data analysis pipelines all add time to the arrival of your results.
Unfortunately, these delays mean that traditional RNA-seq is too slow to replace qPCR in most cases. Waiting weeks for the results to inform common procedures, like tuning screening conditions, optimizing editing efficiency, and tracking differentiation trajectories, is simply impractical, making qPCR the obvious choice for routine experimental work.
Because of this, RNA-seq is rarely used to iteratively guide the evolution of projects. It may be done once at large scale at the beginning or end of a project to establish or test hypotheses, but it’s very rarely used to perform the kind of rapid, iterative refinement loops that are needed to effectively explore complex biology.
This is a shame because the broader perspective provided by RNA-seq could not only transform how experiments are designed but also radically expand the scope of what can be investigated in a study.
How RNA-seq Becomes Practical for Routine Use
While turnaround time is the primary constraint preventing widespread RNA-seq use, it is not the only one. Quote-based procurement, cold-chain logistics, high costs, and analysis complexity each add friction that makes RNA-seq harder to use in iterative experiments.
Removing these barriers is essential to making RNA-seq a practical option for day-to-day use.
Plasmidsaurus, a super cool dino-themed company, provides a 3′ RNA-seq service with a turnaround of three days in the US, UK, and Europe, and around one week elsewhere in the world (because waiting weeks for data is extinct behavior).
The workflow couldn’t be easier:
- Place your order on Plasmidsaurus’ streamlined self-serve online platform. No minimum sample batching or quote required.
- Skip the dry ice. Prepare either lysed cells or purified RNA in SEQguard Dino Preserve so your samples can be shipped at room temperature. (You heard that right, ambient shipping for RNA.)
- Submit your samples at any of Plasmidsaurus’ ultra-local drop-boxes that are located in most global research hubs
- Explore your data the instant it becomes available, with on-platform automated bioinformatics and interactive plots that make analysis a breeze.
So, what makes 3’ end counting different from fragmentation-based RNA-seq?
3′ End Counting: A Simpler RNA-seq for Differential Expression
The strategic mRNA capture scheme and barcoding strategy of 3’ end counting make it inherently more efficient than fragmentation-based RNA-seq, enabling accurate transcript quantification with fewer reads and less downstream correction.
In fragmentation-based RNA-seq, multiple reads per transcript are generated by randomly shearing RNA into fragments before sequencing. After sequencing, transcript counts are calculated by first aligning the fragments to the reference genome and then normalizing them to the expected transcript length.
Because longer transcripts produce more fragments, they can appear more abundant in raw read counts than shorter transcripts of equal abundance if this normalization is not done. This type of miscount is referred to as transcript length bias.
3′ end counting works differently.
Where fragmentation-based RNA-seq needs to sequence all fragments of a transcript to report accurate read counts, the genius of 3’ end counting is that transcript molecules need only be sequenced once for accurate representation in the dataset.
This efficiency means that 3’ end counting needs fewer total reads than fragmentation-based RNA-seq to achieve equivalent sensitivity for differential expression, reducing both sequencing cost and improving throughput.
During reverse transcription, an oligo(dT) primer binds to the poly(A) tail of the mRNA, generating one cDNA molecule per transcript, regardless of how long that mRNA is. Tagmentation is then used to produce a sequenceable fragment at the 3’ end.
PCR amplification is then required to generate enough material for sequencing. In theory, this introduces a risk: some original molecules get copied more times than others by chance. But this problem is entirely removed by the Unique Molecular Identifiers (UMI) that are installed during reverse transcription, so copies of the original molecule can be collapsed to a single count during analysis (otherwise known as deduplication) (Figure 1).
After deduplication, the resulting read counts immediately represent transcript counts, providing a full profile of gene expression in the sample.
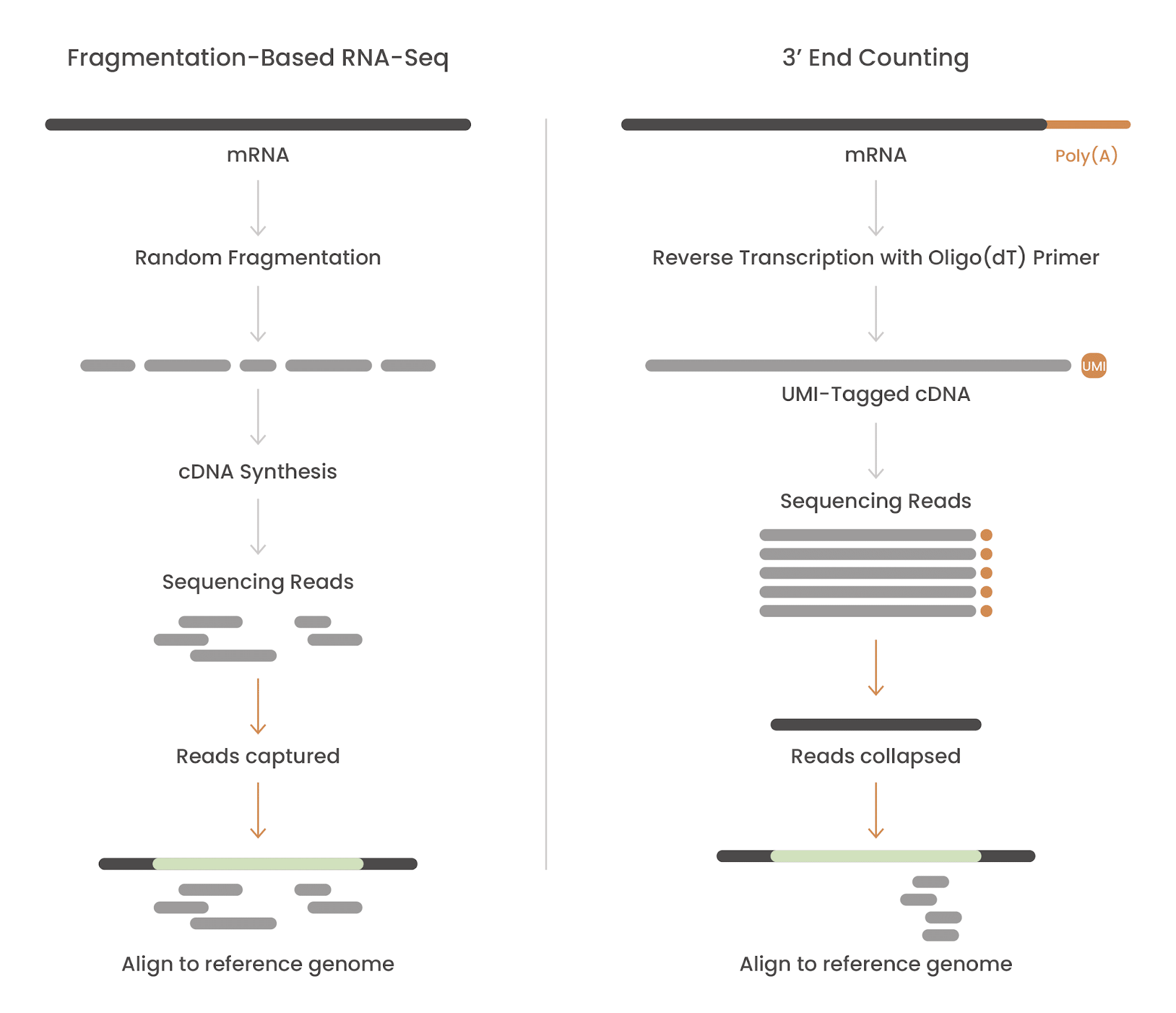
Figure 1. Fragmentation-based RNA-seq vs. 3′ end counting.
While the benefits of 3’ end counting are clear, there are a few considerations to keep in mind. The method is limited to polyadenylated RNA species, which means prokaryotic transcripts lacking poly(A) tails are not compatible with oligo(dT)-based approaches.
Misrepresentation of true transcript levels can be an issue when sample degradation causes mRNA transcripts to lose their poly(A) tails. (Although this can be mitigated with careful sample preparation.)
Finally, because of how 3’ RNA-seq works (it only sequences the end of the gene), it is not able to deliver structure-level insights, such as information about splice isoforms, alternative transcription start sites, and novel transcripts.
For what it’s worth, qPCR also cannot deliver this information, and even fragmentation-based RNA-seq isn’t the best for this. (Where structure-level insights are concerned, long-read full-length cDNA sequencing is the appropriate approach.) However, most gene expression studies don’t need this level of detail.
When Speed Changes Strategy

When experiments take weeks or months, the opportunity cost of testing a more novel hypothesis can be prohibitively high. But when you can get transcriptome-wide data at low cost within days, you’re much freer to explore. You can rapidly test different directions, dynamically scale the scope of your experiments, and chase new insights you find in the data (Figure 2).
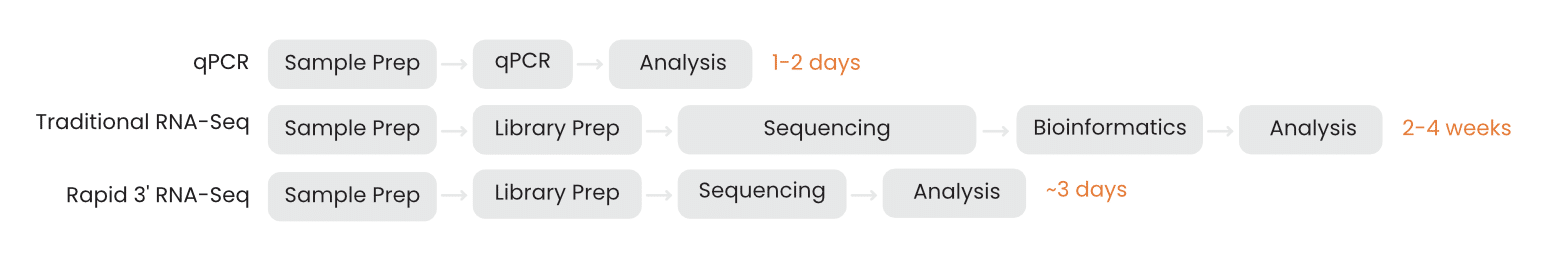
Figure 2. Turnaround time shapes experimental strategy.
This reduced RNA-seq timeline means you can use NGS much earlier in your study, shifting your research from confirming a hypothesis to discovering what’s actually happening. For any type of study, rapid RNA-seq means that you can incorporate it into your workflow for faster feedback loops and optimization, allowing you to make more informed decisions every step of the way. Here are a few examples:
In drug response profiling, understanding what is happening at the genome-wide level can help you screen candidates more efficiently and improve the chances of successful clinical translatability.
With qPCR, you could detect the change to a specific pathway and prove or disprove your specific hypothesis. But with RNA-seq, you can profile the whole transcriptome and see everything that changed in response to the drug—including compensatory responses and pathway crosstalk that would otherwise remain undetected until a later experimental stage, when correcting course is more costly.
Speed is also helpful if you’re looking at transcript-wide effects of different guide RNAs in CRISPR studies. If you used a targeted qPCR panel to characterize guide RNA effects, you risk missing off-target transcriptional changes. With a fast RNA-seq turnaround, you can compare multiple conditions in parallel before committing to a lead candidate.
For differentiation and transcription factor workflows, you can collect transcriptome-wide data for each optimization round with RNA-seq, which gives you more information to work with in the next step. Plus, this ability to run more iterations in the same amount of time means you can complete whole optimization cycles much faster.
Choosing qPCR or RNA-seq for Your Experiment
Ultimately, the decision between qPCR, traditional RNA-seq, and 3’-end counting depends on your experimental goals.
If the goal is to assess the differential expression of a small, pre-defined gene set, qPCR can still be a practical choice. It is fast, keeps pace with day-to-day lab work, and requires no complex downstream analysis. However, qPCR is primarily used because of its accessibility rather than its technical capabilities.
RNA-seq becomes more attractive as the number of target genes in a gene expression study increases (or where the cost of missing something unexpected is high). In most drug screens, CRISPR validations, and differentiation studies, the key question is not which few genes changed, but what changed across the system. These types of investigations would not just benefit from high-throughput sequencing; it is effectively essential to do so. However, long lead times and complicated analysis have largely limited the broader use of traditional RNA-seq.
Plasmidsaurus’s 3’ end counting RNA-seq service has overcome all of these logistical challenges, making RNA-seq more accessible than ever before. Now that transcriptome-wide profiling fits within the same turnaround time as a targeted qPCR assay, the decision to perform DGE studies with 3’ end counting RNA-seq couldn’t be easier. Why? Because choosing between speed and scale is a compromise that may leave your experiments moving at a pace better suited to the Jurassic era than to modern research.
Want to learn more? Check out Plasmidsaurus’ website for more information, or watch Expression Data in Days, Not Weeks: Accelerating Biology with Plasmidsaurus 3′ RNA-Seq on YouTube today.
References:
- Oshlack A, Wakefield MJ. Transcript length bias in RNA-seq data confounds systems biology. Biol Direct. 2009;4:14. https://doi.org/10.1186/1745-6150-4-14
- RNA-seq, www.plasmidsaurus.com/rna
- Expression Data in Days, Not Weeks: Accelerating Biology with Plasmidsaurus 3′ RNA-Seq, youtube.com