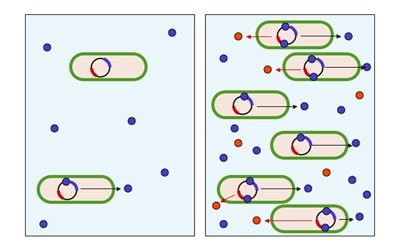
RNA interference (RNAi) may have originated as a defense mechanism to protect cells against foreign genes introduced by viruses. This concept has since been put to use to create a powerful experimental tool for investigating gene function in organisms. Small-interfering RNA (siRNA) libraries for investigating genome-wide function can be produced by chemical synthesis of probes or vector-based small-hairpin RNA (shRNA) libraries.
Here I will describe the technology and parameters that go into designing these siRNA probes.
What is RNA interference?
RNA interference (RNAi) is a conserved mechanism found in most eukaryotes. RNAi uses short double-stranded RNAs (dsRNAs), usually 21-24 nucleotides long, and RNA-induced Silencing Complex (RISC) machinery to silence expression of genes with complementary sequences to the dsRNA. In cells, long dsRNAs, which normally originate from viruses, are degraded by a protein named Dicer, an RNase III ribonuclease family enzyme that is a component of RNA interference pathway. Dicer degrades long dsRNAs to 21-24 nucleotide-long double-stranded small interfering RNAs (siRNAs). These siRNAs are then incorporated into the RISC machinery, which is made up of a number of proteins, including Dicer, TAR RNA-binding protein (TRBP) and Argonaute 2 (AGO2).
While both strands of siRNA could potentially be used for silencing, only one strand in the duplex, referred to as the guide strand or silencing strand, is incorporated into RISC, where it assembles with AGO2. Argonaute proteins contain two RNA binding domains: a PAZ domain that interacts with the 3’ end of miRNA, and a PIWI domain that resembles ribonuclease-H and is responsible for the endonucleolytic degrading activity. Typically, the strand with a lower base pairing stability of the 2-4 bases at the 5’ end of the duplex is preferentially associated with RISC, and thus becomes the active guide strand. Guided to its mRNA target by the guide siRNA strand, the RISC complex then degrades the mRNA target, effectively “silencing” the RNA.
Choose a free resource to help you move forward
CHEAT SHEET
SDS-PAGE Protocol Cheat Sheet
EBOOK
Free Guide to Protein Expression
Tips for designing siRNA probes
It is not surprising that siRNAs represent promising reverse genetic tools for answering fundamental questions about gene function, performing genome-wide discovery studies, and developing potential therapeutic agents to targeting pathways associated with pathogenesis. The power of this technology, however, is limited by the efficacy of siRNA probes and their ability to specifically silence the target gene. The design of siRNA probes, therefore, represents an essential step in increasing the potential application of this technology not only for small-scale screens but also for high-throughput RNAi screening initiatives in mammalian systems. Here are some tips for how to design effective siRNA probes:
- Sequence location: The ability of an siRNA to silence gene expression depends on the sequence complementarity between the silencing RNA and the target. However, the exact location within the target also plays a critical role in determining silencing efficiency. Therefore, it is critical to first confirm the presence of complementary sequence in the transcript variant, since alternative splicing of the target gene may result in omission of certain sequences complementary to the siRNA probe.
- Chemical modifications: Chemical modifications of the siRNA probe may enhance its ability to bind to and silence the target transcript. Certain modifications can affect the half-life of the siRNA by making it more susceptible to nuclease degradation, or they can improve cellular uptake and decrease off-target effects of the probe. However, these modifications must not jeopardize the integrity of the siRNA. For example, all siRNAs have a phosphate group at the 5’ end, so it is important not to introduce modifications that would block the 5’ end of the molecule.
- Thermal Stability: The siRNA duplex must separate to allow one of its strands to be loaded into the RISC machinery. The 3’ end overhang is important in siRNA function and RISC loading. Introducing modified bases at the 3’ end can be used to influence the duplex stability and manipulate which strand will be loaded into RISC. It has been shown that sequences with higher thermal stability at the 5’ end of the sense strand compared to that at the 3’ end are better able to direct entry of the minus (targeting) strand of an siRNA into the RISC complex, and so promote more effective RNAi while presumably decreasing the likelihood of off-target silencing. Additionally, it is possible to introduce sequence asymmetry to the duplex via mismatches, in order to influence which strand is preferentially loaded into the RISC complex.
- GC content: GC content has a strong impact on siRNA silencing efficiency. siRNA sequences should not contain homopolymeric sequences of four or more identical nucleotides, runs of nine or more GC residues, and should have an optimal GC content of between 30 to 50%.
Using siRNAs in vitro and in vivo
Synthetic siRNAs have been used extensively as a research tool for investigating and validating gene function and functional pathways, and have also been used as therapeutic agents in clinical applications. Although siRNAs provide high knockdown efficiency, the knockdown is transient, thus limiting the use of siRNAs to short-term experiments. While siRNA specificity can be validated in vitro, several points need to be taken into consideration for in vivo integration: limiting off-target effect (which can arise when the siRNA has a sequence that can pair and reduce the expression of multiple genes at a time), determining the optimal route of administration and delivery method, and facilitating cellular uptake while minimizing the immune response. To circumvent the transient nature of siRNA regulation, lentiviral systems have been used as part of the gene therapy approach to allow stable and permanent integration of siRNA sequences into the genome.
RNA interference is an endogenous gene-silencing pathway in living cells that has become a valuable research tool for high-throughput genomic studies as well as therapeutic applications. In order to deliver maximum efficiency with minimal off-target effect, several parameters mentioned above must be taken into account when designing synthetic siRNA probes.
What advice do you have for designing effective siRNAs?
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.