We all know the impact fluorescent proteins have had in advancing cell biology. Although fluorescent proteins have revolutionized the field, they aren’t perfect and like all things research, they have their limitations. If you’re looking for a genetic tool with superior fluorescent properties, or one that allows you to introduce a variety of labels into your system, then you should look into self-labeling proteins (SLPs).
How Self-Labeling Proteins Work
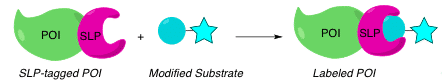
SLP is a technology based on the formation of a specific, covalent bond between a labeling molecule and a fusion tag of a protein of interest (POI). To use this tool, you’ll first clone and express your POI as an SLP fusion. Then, you’ll tag the SLP with a modified substrate that is linked to a label.
The label can be virtually any molecule of your choice. Pick a fluorescent dye, a capture tag, or whatever probe you need, based on what you want to study. And as if that isn’t enough to get you excited, the protein labeling reaction is typically fast and quantitative, making it ideal for a variety of in vivo and in vitro applications.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
SDS-PAGE Protocol Cheat Sheet
EBOOK
Free Guide to Protein Expression
Let’s go over the steps involved in running an SLP experiment:
Step 1: Genetic Fusion
An SLP is a protein tag expressed as a fusion of a POI. Generally, you may clone the gene encoding your POI into the vector on either side of the tag without affecting the function of your protein. The most widely used commercial SLP systems are Promega’s HaloTag® and New England Biolabs’ SNAP-tag® and CLIP-tagTM. Both suppliers offer a variety of SLP kits with cloning vectors for expression in mammalian and bacterial systems to get you started.
Step 2: Covalent Labeling
Most often, SLPs are an engineered version of enzymes that react covalently with their substrate. The substrate is modified to a variety of synthetic derivatives carrying a labeling molecule. When you incubate your SLP-tagged POI with the substrate of your choice, a stable bond between the SLP and the substrate forms, resulting in your POI becoming labeled as desired.
Before deciding which SLP method to use, let’s take a closer look at the technology behind HaloTag, SNAP-tag, and CLIP-tag.
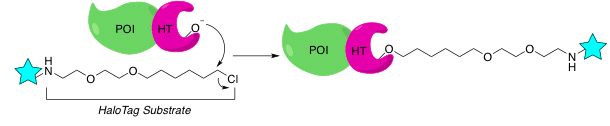
HaloTag
The HaloTag protein is a mutant of a bacterial dehalogenase. This enzyme removes halogens from aliphatic hydrocarbon molecules through a nucleophilic aspartate residue. While the wild-type enzyme is able to regenerate its catalytic site for further dehalogenation, engineering of the HaloTag protein allows it to form an irreversible covalent bond with its substrate.1 This type of linkage allows the protein to form a stable bond with any labeling chloroalkane ligand of your choice, a feature you will find useful for downstream applications.
Note: The absence of the native protein in eukaryotic cells and most prokaryotes, including E. coli, allows you to use a variety of expression hosts to achieve specific labeling of your POI.
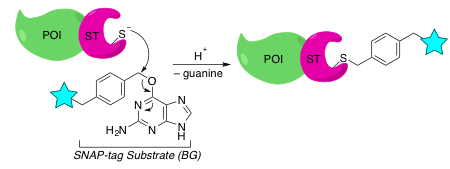
SNAP-Tag
The SNAP-tag technology is based on human O6-alkylguanine-DNA-alkyltransferase (hATG). This DNA repair enzyme transfers the alkyl group of mutagenic O6-alkylguanine-DNA to one of its cysteine residues to release guanine. Taking advantage of this irreversible process, the SNAP-tag protein was engineered from hATG to specifically recognize and react with O6-benzylguanine (BG), a guanine conjugated to a label.2
During covalent labeling, the catalytic cysteine of the SNAP-tag forms a stable bond with BG’s labeling group, which in turn gives off (repairs) guanine. This reaction is highly specific, and the chemical inertness of BG also adds peace of mind when it comes to off-target labeling.
Note: You may use any expression host, and the wide variety of labeling substrates allows you to label at the cell surface and virtually anywhere within the cell.
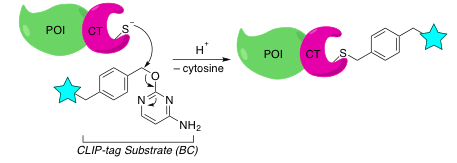
CLIP-Tag
If you’re looking to do dual labeling, then the CLIP-tag is for you. The CLIP-tag was developed to complement the SNAP-tag so that you can apply both types of fusions in one experiment. The CLIP-tag protein is also a derivative of hATG, so it shares the features of its sibling, the SNAP-tag protein. This SLP was generated to specifically recognize and react with O2-benzylcytosine (BC), a cytosine conjugated to a label.3
As you may have guessed, the principle behind the labeling reaction is similar to the SNAP-tag technology. The basic difference is that you’ll use a BC derivative instead of BG to label your CLIP-fused POI. The catalytic cysteine of the CLIP-tag reacts with BC to repair cytosine, forming a stable bond with the labeling group.
Note: By using both SNAP- and CLIP-tag technologies you will have the opportunity to simultaneously label two fusion proteins with two different probes.
Scope and Applications for SLPs
Most commonly, fluorescent molecules are linked to the substrate to enable cellular imaging applications, such as protein localization and function analysis.4 Although this concept is similar to that of auto-fluorescent protein tags, SLPs have several advantages over fluorescent proteins, including:
- A broader choice of fluorescent dyes
- Higher fluorescence intensity and photostability
- Fluorescence initiated when the label is added (temporal control)
- A single construct is sufficient to label with different fluorophores (multicolor imaging)
- Fluorescence remains strong in both live and fixed cells
Now, this doesn’t mean you should give up on your favorite fluorescent protein. You may pair the SLP technology with fluorescent protein-tags to run a really cool, dual labeling experiment to study protein-protein interactions, for instance.
If you’re yet not convinced of the power provided by SLPs, then you may be interested in knowing that this technology isn’t limited to glowing substrates. Other suitable labels include biotin, magnetic beads, and gold particles to enable affinity binding or protein immobilization.
Preparing Your Own Substrate
What if you can’t find a substrate with the label you need, you ask? Well, you can custom-make it from commercial SLP building blocks. For example, buy an SLP substrate with a reactive functionality, such as a primary amine, and link it to your desired label containing an activated ester functional group. And if that label doesn’t work, or you want to try something else, then link the same building block to any other label you want.
Fortunately, there is a plethora of commercially available labeling compounds you can choose from to prepare your own substrate. But if you’re looking for something completely new and feel like being creative, then collaborate with a synthetic chemist to develop a novel labeling substrate. Why not, right? Interdisciplinary collaboration is always fun and you get to learn so much!
The choice of label and application is yours, and the possibilities are up to your imagination! (Well, yes, and up to funding availability, but that’s a topic for another day!)
Like the sound of that? Perhaps SLP is the technology your research needs. What type of labels would you like to use? Let us know in the comments.
Upload More Knowledge to Your Brain
- Los GV, Darzins A, Karassina N, Zimprich C, Learish R, McDougall MG, Encell LP, Friedman-Ohana R, Wood M, Vidurgiris G, et al. (2005) HaloTagTM interchangeable labeling technology for cell imaging, protein capture and immobilization. Promega Cell Notes. 11: 2–6.
- Keppler A, Gendreizig S, Gronemeyer T, Pick H, Vogel H, Johnsson K (2003) A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21: 86–89.
- Gautier A, Juillerat A, Heinis C, Corrêa, Jr. IR, Kindermann M, Beaufils F, Johnsson K (2008) An Engineered Protein Tag for Multiprotein Labeling in Living Cells. Chem. Biol. 15: 128–136.
- Provost CR and Sun L (2010) Fluorescent Labeling of COS-7 Expressing SNAP-tag Fusion Proteins for Live Cell Imaging. J. Visualized Exp. 39: 1876
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
SDS-PAGE Protocol Cheat Sheet
EBOOK
Free Guide to Protein Expression