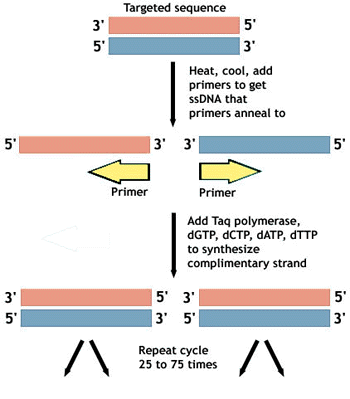
Applying molecular techniques to unicellar organisms leads to many questions…
- Did my electroporation work?
- Is my vector inside my competent cell?
- Do I have contamination in my liquid culture?
- Is this the correct bacterial strain the neighboring lab promised me it is?
- Did the guy from the other side of the world send me the mutant I asked for?
All these questions can be answered by going through the slow, routine pipeline process in a molecular biology laboratory: Culture cells, harvest them, extract DNA and PCR.
Or guess what?! You can directly perform PCR using the colonies on the petri dish that the post-doc left on your bench. Some people refer to it as “Colony PCR” while others call it, “fast boiling”; whatever the name, it is a time saver for sure.
So next time you wonder if you need to spend your weekend screening the 200 colonies the post-doc asked you to check, the answer is definitely NO.
Put this article into practice
Choose a free resource to help you move forward
DOWNLOAD
Blood Collection Tube Chart
CHEAT SHEET
Western Blot Cheat Sheet
How to?
First, make sure you know what you are dealing with. The process can vary (not much) for gram+ and gram– bacteria, microalgae or even bacteriophages.
Second, set up your PCR.
- Grab some PCR tubes and add 20μl ddH2</sub
- Using your pipette and a tip, barely touch the colony or the petri dish in which the microorganism is spread and pipette it into the PCR tube
- Put the tube in your PCR machine and us the following program: 95°C for 15 minutes, 10°C for 15 minutes (for preservation or finishing your coffee)
Third, take 1&mul from the first PCR reaction and use it as your DNA template in a standard PCR for your gene of interest.
Fourth, take a photo of your band and feel awesome!
Of course this technique can vary depending on your starting colony. If you are dealing with microalgae (in which probably you will now understand that you are going to save maybe a week because microalgae can take up to 2 weeks to reach sufficient yield for DNA extraction) add 1&mul of 1N NaOH in your mix and increase the boiling time to 30 minutes. If you are dealing with a phage, things can get trickier because some of them require a protease treatment to release the DNA.
Troubleshooting and tips
- Always have a positive control to make sure the reaction worked.
- You need only a very little amount of LIVE cells, so make sure you barely touch the tip to the plate. The cells can contain inhibitors to the PCR reaction. If you think your mixture is too dense, make a 1/10 dilution after boiling and PCR that as well. Avoid taking agarose from your plate, this can be a major inhibitor of your PCR reaction.
- Store your DNA template at -20°C for up to 7 days. If you use it more than once, DNAases from the dead cells can degrade the DNA, while you are calculating your PCR reagents. So start again from the colonies.
- Boil cells from fresh colonies only. Do not use old colonies left in fridge for 6 months. Do not forget, more cells in a colony are DEAD, while fresh cells contain the maximum amount of DNA as they are in a replication frenzy mode.
- You can perform the same technique starting with liquid cultures depending the density of the culture.
- Spin down your PCR tubes after boiling to separate debris cells from nucleic acid.
- If your supervisor disagrees with this technique, remind him/her HOW MUCH MONEY you saved in DNA extraction reagents and that if this technique is adopted by the lab, in 10 years he/she will save a lot of money.
And remember: a positive band will definitely indicate a positive result (assuming you do not have a contamination), but a negative result could be a false negative (you might not have recovered enough DNA, or you have too many inhibitors in your reaction).
Good luck and I wish you strong bands.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
DOWNLOAD
Blood Collection Tube Chart
POSTER
Cell Culture Posters