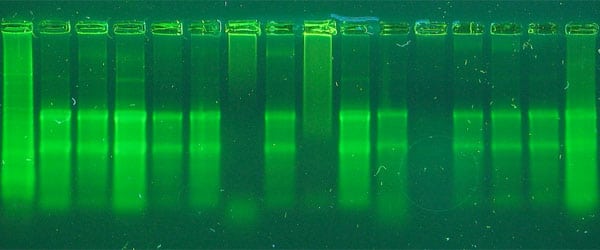
One of my favorite things to do with a student the first time they work with DNA plasmid preps is to have them run plasmid DNA on agarose gels. I get them to run two samples: uncut plasmid DNA, and plasmid DNA that has been linearized with a restriction enzyme.
I love to have them try and figure out the banding pattern of uncut plasmid DNA (why do you see 2–3 bands?) versus the single band of linearized DNA (why do all bands convert to 1 band when linearized?).
I like this exercise because understanding the forms on the gel requires an understanding of the nature of DNA.
In this article, I will focus on the four most common species of uncut plasmid DNA observed on a gel and how to recognize them.
And in my next article, I’ll cover how to increase the recovery of the desired supercoiled species.
One Molecule, Many Forms: Why Uncut Plasmid DNA on Agarose Gel Has 3 Bands
When uncut plasmid DNA is isolated and run on an agarose gel, you may observe two, three, or even four or more bands. Hopefully, the majority of your isolated DNA will be supercoiled, but other forms can also crop up. How these forms will show up on an agarose gel (in terms of relative migration speeds) is shown in the diagram below:
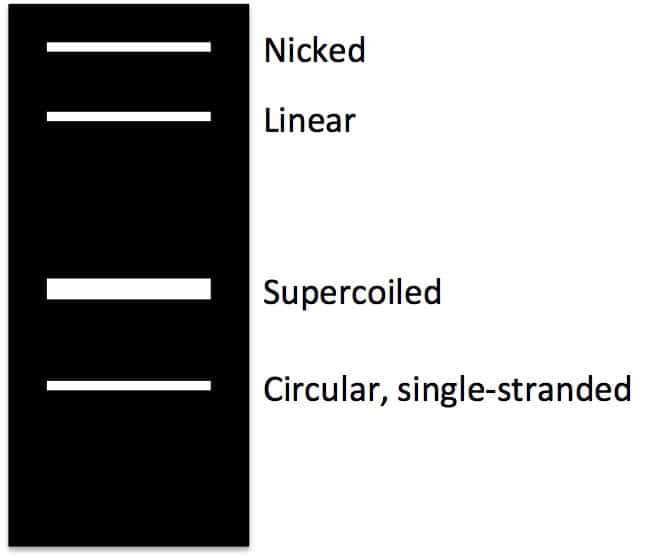
And here I’ll take you through where each form comes from.
Supercoiled Plasmid
Supercoiled DNA is the native DNA conformation found in vivo and occurs when extra twists are introduced into the double helix strand. People often compare the forms of DNA to rubber bands or telephone cords (I know some of you must still remember phones with cords!). If you over twist a rubber band or telephone cord, coils stack up on one another, introducing tension.
In the case of DNA plasmid preps, this superhelical tension cannot be relieved because the ends of the plasmid are joined together. Supercoiled DNA migrates faster than predicted in an agarose gel due to its conformation. Supercoiled DNA is the desired species when isolating plasmid DNA.
Nicked, Relaxed, or Circular Plasmid
DNA found in the supercoiled form is not easily accessed by the replication machinery. During replication, cellular topoisomerases nick one strand of the DNA helix and relax the superhelical tension, thus allowing polymerases to gain access to the DNA. Using the rubber band analogy, nicked circle DNA is the rubber band without any twists introduced. This large floppy circle is the slowest migrating form in an agarose gel.
Linear Plasmid
Linearized DNA occurs when the DNA helix is cut in both strands at the same place. Linear DNA generally migrates between the nicked circle and the supercoiled forms. However, it may also migrate the same distance as the nicked circle – it migrates as predicted by the length of the DNA (as compared to the molecular weight markers).
You can identify the linear DNA form on an agarose gel by comparing uncut plasmid DNA with a sample of the plasmid that has been linearized using a restriction enzyme. If you get linear DNA when you are hoping for supercoiled (e.g. after DNA plasmid preps), it is due to nuclease contamination or harsh treatment during purification.
Circular, Single-Stranded Plasmid
During alkaline lysis plasmid preps, plasmids are denatured because the hydrogen bonds are disrupted by the alkaline conditions. But the covalently closed circular strands remain intact and topologically constrained and when the pH is returned to neutral the hydrogen bonds reform and the supercoiled DNA is re-formed.
However, if the alkaline lysis step is overly harsh (e.g. it is incubated for too long), the DNA can become permanently denatured and give you useless single-stranded closed circles that migrate ahead of all of the other forms of the plasmid in a gel.
Successful DNA Plasmid Preps
Although DNA plasmid preps can return multiple forms of DNA, there is only one kind you want for successful cloning and transfection: supercoiled. Make sure you know how to increase your recovery of good quality supercoiled DNA. Did this help you understand why you get three bands when running plasmid DNA on agarose gels? Do you have any other plasmid prep tips? We’d love to hear in the comments.
Originally published October 8, 2014. Reviewed and republished April 2021.