I often wonder why it is that molecular biology researchers stubbornly refuse to change 40-year old methods that, while work, are not as good as newer, faster and cheaper methods out there.
I suppose rational scientists often have irrational superstitions.
One example of an old method that could be improved is the growth media used for plasmid preparation.
The majority of us, throughout our university careers, have used either SOC, LB or TB, for recombinant plasmid propagation, typically in E. coli. LB or Luria-Bertani broth has been in use for almost 60 years or thereabouts, while SOC has certainly been in use for 2 decades.
Pimp Your media and Save Some Money!
But by adding in a few more ingredients or being more economical on others (especially yeast extract and tryptone) you could get a higher plasmid yield, quicker and with less money.
To counter the naysayers, nobody wants to make very complex media with 15 ingredients requiring filter sterilisation, as this obviously defeats the object of economy of time and budget. Indeed, there are trade-offs between optimising for biomass, plasmid yield, quality, stability and cost with the difference between protein production and plasmid production, since plasmid production requires only cell growth, division, and plasmid stability.
The good news is that Michael Danquah and Gareth Forde from Monash University down-under have devised a stoichiometrically optimised medium for plasmid production. PDM, supposedly yields under the conditions they tested, twice the amount of plasmid in both volumetric and specific yields compared to TB, and LB is left in the dust. Better yet, because it uses less tryptone and yeast extract, the cost per mg of DNA is roughly one-quarter of the cost of LB.
The recipes for LB, TB, SOC and PDM are shown below. If you decide to break with tradition and give PDM a go, be sure to tell us how it goes.
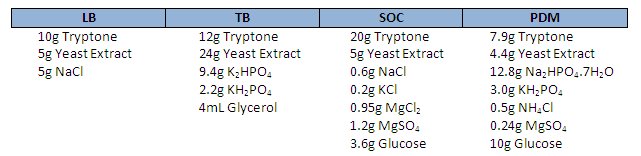
Note: Autoclave glucose, KH2PO4 and Na2HPO4 separately – if you want to know why check out Doesn’t Play Well with Others – The Chemistry of the Autoclave.
Originally Published 28 April 2008, Updated and Republished 19 December 2014.