Here’s an all-too-often repeated scene in the lab: First thing in the morning, you approach the 37°C incubator with trepidation, open it and through one half-open eye you take a look at the LB plate that you spread your ligation-reaction-transformed E.coli aliquot onto.
Looks good – thousands of colonies.
Emboldened, you take your “no ligation control” plate.
Looks bad – thousands of colonies.
Argh! Somehow, uncut or religated vector made it through to the transformation, and your clones contain nothing but empty vector.
Before this happens to you again, consider these two tips for preventing empty vector from getting into the final transformation step of your cloning experiment.
1) Use less Vector
A common rule of thumb is to use 100 ng of vector in ligation re
actions, but this is too much. Most ligation protocols actually suggest 5-15 fmol, which is much less than 100 ng for your typical size vector. Adding in too much vector stacks the ligation reaction favor of empty recombinants
Ligation reactions are quite efficient so the important thing is to present the ligase with just enough vector to give a high chance of the vector and insert meeting, without over-doing it and increasing the chances of a self-ligation or concatenation reaction.
2) Use a killer cut
Using a killer cut is a superb trick because it efficiently eliminates empty recombinants right at the end of the cloning process, just before transformation. To make a killer cut, you need to have a restriction site that linearises the original vector but does not not cut the recombinant.
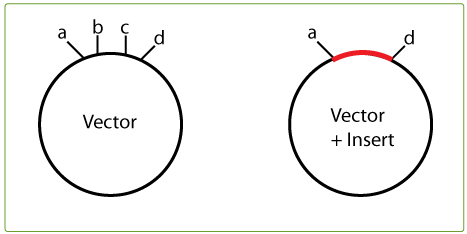
To find your killer site, take a look in the multiple cloning site of the original vector and note the restriction sites that are between the two cloning-in sites you are using; in the example on the right, this would be restriction sites b and c.
Then check whether b or c cut in the insert. If, for example, B does NOT cut in the insert, it can be used as a killer cut since it will linearise the vector, but not the desired construct (vector + insert).
So at the end of the ligation, simply add 1/100th volume of the killer cut enzyme, incubate at the optimal temperature for 10 minutes and it will linearise, and therefore prevent the transformation of, most of the empty vector recombinants. Say goodbye to the pesky colonies on the control plate.
Although the restriction enzyme is not in its optimal buffer, it should easily be able to cope with the tiny amount of plasmid in the ligation reaction even if it is performing sub-optimally.
If you use these approaches, or if you try them after reading them, let us know your experiences and any additional tips you might have.
(This article is based on a tip emailed to me by Julie Wolf of UMBC, Maryland. Thanks Julie!)