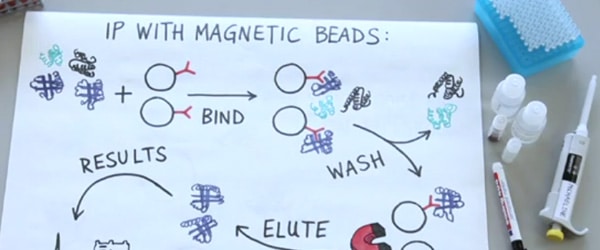
Do you want to immunoprecipitate (IP) a protein with a molecular weight that is anywhere near 55 kDa or 25 kDa? Then you have an irritating problem to deal with: antibody co-elution. But don’t panic, we have six strategies for dealing with your new problem.
The Problem:
Typically, the IP antibody is bound to Protein A-coupled beads (or Protein G beads). Because this method of attachment is non-covalent, when you elute the target protein a large amount of unwelcome antibody can be eluted too. The co-eluted antibody runs on a denaturing gel at around 55 kDa (heavy chain) and 25 kDa (light chain), and these contaminating bands can interfere with downstream analysis in a variety of annoying ways.
The problem isn’t solved by using standard western blotting to detect the target, because you usually need to use the same primary antibody for detection that you used for the IP. This means that the secondary antibody will recognize two overlapping signals: the functional primary antibody that is bound to your target bands, as well as the denatured heavy and light chain.
Choosing a Strategy:
- If the co-eluted antibody interferes with mass-spec analysis, or obscures bands on a stained gel, or causes band deformation:
Then you need to prevent the antibody from co-eluting. Try strategies 1, 2 and/or 3.
Put this article into practice
Choose a free resource to help you move forward
CHEAT SHEET
Western Blot Cheat Sheet
EBOOK
Free Guide to Protein Expression
- If the co-eluted antibody obscures bands on a western blot:
Then you don’t necessarily need to stop the antibody from co-eluting, you just need to avoid detecting it. Try strategies 4, 5 or 6.
Strategy 1: Crosslink the antibody to Protein A/G beads
One way to keep the antibody in its place is to covalently crosslink it to the beads. The antibody is first captured by Protein A or G beads and then treated with a reagent like dimethyl pimelimidate (DMP) or one of the many commercial crosslinking kits. Too much crosslinking can decrease the antigen-binding activity of the antibody, so you might need to optimize the crosslinking conditions.
Strategy 2: Couple the antibody directly to the beads
It is also possible to skip the Protein A/G binding step by directly coupling the antibody to chemically activated beads. Rather than treating the beads yourself, it is simplest to buy pre-activated beads such as cyanogen bromide-activated agarose, agarose N-hydroxysuccinimide esters, or tosyl-activated magnetic beads. Direct coupling is usually applied to purified antibodies, because there is no purifying Protein A/G binding step and the activated beads can react with contaminating proteins and amine-containing compounds. Again, antigen-binding activity can be decreased by ‘overcoupling’, so you might need to optimize the coupling conditions.
Strategy 3: Use milder elution conditions
For some antibody-antigen combinations, elution conditions can be found that are strong enough to release the target protein without releasing the antibody. In some cases, you may be able to get away with simply leaving out the reducing or boiling step from a stringent elution with SDS-PAGE sample buffer. In most cases, however, choosing the ideal elution conditions will require some optimization, but you could start by testing the effect of pH, ionic strength or the presence of denaturants. Glycine at pH 2-3 is a very common eluant, and titration of different ratios of ionic and non-ionic detergents may also work.1 Unfortunately, mild elution conditions will usually give lower final yields than boiling in sample buffer.
Strategy 4: Use radioisotopes
If the problem is just that the co-eluted antibody interferes with your bands in a western blot then a traditional remedy is to radiolabel the cells before making the cell lysates. The IP is performed as usual and the protein gel is visualized as an autoradiograph. This means that only proteins from your cell lysate will be detected in the elution mixture. The upside is that isotopic detection is sensitive. The downside is that you have to deal with radioisotopes.
Strategy 5: Use a different antibody for the western
A simple option is to use an antibody from a different species for the western detection step, since the secondary antibody then won’t recognize the contaminating antibody.
A less-common variant of this strategy is to skip the secondary antibody altogether and use primary antibodies coupled to a secondary detection molecule (e.g. Horseradish Peroxidase). Unfortunately, such primary antibodies are not always commercially available and direct detection methods are usually less sensitive than secondary detection.
Strategy 6: Use labelled Protein A/G for secondary detection
An ingenious fix for all those times when you don’t have access to multiple kinds of primary antibody is to replace the secondary antibody with Protein A/G coupled to a secondary detection molecule2. Protein A/G won’t bind the denatured antibody from your IPs, but will bind to the native antibody used in the western blot, rendering the contaminant bands invisible.
The Control
Don’t forget the essential control for any of these strategies: a mock IP. Add lysis buffer to the IP instead of cell lysate, and perform the same steps as usual in parallel with your sample. This allows you to quantify the amount of antibody that is co-eluted.
We’d love to hear how you get around this problem, feel free to share your tips and tricks in the comments!
References
1Antrobus, R, Borner, GHH (2011). PLoS ONE 6(3): e18218
2Lala, A, Haynes, SR and Gorospea M (2005). Molecular and Cellular Probes 19(6) pp 385-388.
You made it to the end—nice work! If you’re the kind of scientist who likes figuring things out without wasting half a day on trial and error, you’ll love our newsletter. Get 3 quick reads a week, packed with hard-won lab wisdom. Join FREE here.
Put this article into practice
Choose a free resource to help you move forward
DOWNLOAD
Western Blot Troubleshooting Card
EBOOK
Free Guide to Protein Expression