Sometimes only a small subset of a cell population will show apoptotic features making flow cytometry an excellent way to identify and quantify them. A previous Bitesize Bio article showed how flow cytometry can detect apoptotic hallmarks. More than 30 different dyes can be used to detect apoptosis. It is also true to say that they pick up single apoptosis features rather than the whole biochemical process.
“The needs of the many outweigh the needs of the few”
All multicellular organisms must balance cell proliferation and cell death (apoptosis). Cells are often accidentally lost through trauma but there are situations where cells choose to die. A good example of the latter is deciduous trees shedding leaves.
Several biochemical pathways allow cell death—they are often lumped together in the term apoptosis but, unless specifically defined, apoptosis also includes autophagy, necroptosis and paraptosis (regulated necrosis and an alternate form of cell death, respectively). The term apoptosis follows from work in the early 1970s, when unusual cells were identified in tissue sections and is the process we will consider detecting here.
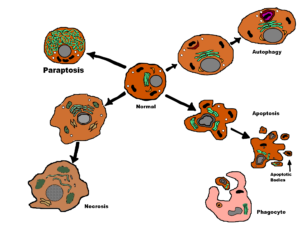
Apoptosis is a tightly regulated biochemical process involving the activation of specific pathways, mainly centred around a family of enzymes called caspases. Cellular apoptosis can be morphologically identified by cell shrinkage, marginalisation of chromatin, and the breakup of cells into apoptotic bodies.
Apoptosis may be induced by external environmental changes like activation by small molecules like Tumour Necrosis Factor (TNF). This is the extrinsic or death receptor pathway.
In the intrinsic pathway, apoptosis induction comes from within the cell. Cells monitor their health. If things get too bad (for example through excessive DNA damage), then cells choose to die via release of cytochrome c from the mitochondria.
Both the extrinsic and intrinsic pathways set in motion a chain of reaction packaging the cell into fragments that are removed by phagocytosis.
How Can We Detect Apoptosis by Flow Cytometry?
We can broadly detect four parts of the apoptotic pathway:
- Changes to organelles, notably the mitochondria. We can measure mitochondrial membrane potential with dyes such as TMRE and JC-1. When a cell enters apoptosis, the proton gradient across the mitochondrial membrane dissipates. The loss of this gradient causes reduced fluorescence from these dyes. You can also track the release of cytochrome c via a fluorescently-tagged antibody method.
- Activation of specific caspases, especially caspase 3, also known as the “executioner of apoptosis.” Several antibodies exist that are specific to the active form of this enzyme, so active caspase 3 will only be detected in apoptotic cells.
- Changes to the plasma membrane. One of the classical flow cytometric methods to detect apoptosis is using annexin V binding to phosphatidylserine residues normally located within the plasma membrane. Phosphotidylserine residues are externalised during apoptosis, so only cells that have decided to die will be detected by annexin V binding.
- Changes to nuclear DNA. As apoptosis progresses, DNA becomes fragmented. You can detect the fragmented DNA by an end-labelling technique or simply treating the cells with detergent to extract the DNA fragments and looking at DNA content. This is the widely used “Sub-G1 method” where apoptotic cells show DNA content below normal G1 cells. However, there are some caveats of this method discussed in an earlier article.
All these methods need to be interpreted carefully, because other cells, not only apoptotic cells, can exhibit any one of these particular features. However, if you look at several methods and combine this with morphological or molecular approaches you can be more confident about detecting the apoptotic process.
So, Which Technique Should I Use?
Because there are a lot of available techniques, an obvious question is which method should we use? Well, the answer will depend on several factors:
- The cell type being used. Are you looking at cells grown in suspension, as monolayers, or primary tissue. Each requires a different preparation and approach. Protocols looking at membrane changes, for example, may not be appropriate on adherent cells.
- The nature of the inducer. We may know which pathway is activated by a particular drug that could influence the method used. But if we are not sure how apoptosis is proceeding, then we may need to try several methods.
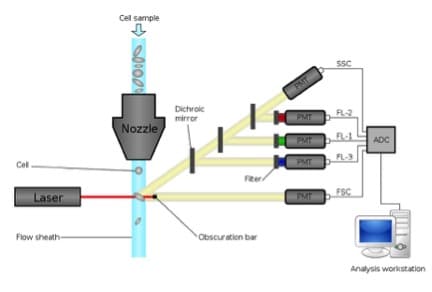
- The cytometer and expertise available. For all flow cytometry experiments, we need to know which lasers are in the cytometer, because the fluorochromes we can use depend on the wavelength of the lasers. Also, we need to know which optical filters are available to detect the emitted fluorescence.
- The cost of the method. We all are acutely aware of lab budgets and some reagents are more expensive than others, so check cost before planning that 200 tube experiment!
Of course we can also combine methods—this is one of the great things about flow cytometry. So, for example, we can look at annexin V binding and changes in mitochondrial membrane potential, or release of cytochrome c and DNA content. The combination will depend on what you know about the system, the compatibility of the fluorochromes, and what information is being sought. But with even a simple flow cytometer there are a plethora of choices!