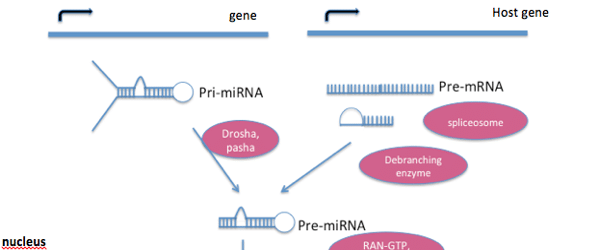
Kary Mullis invented polymerase chain reaction (PCR) in 1985 creating a revolution in molecular biology techniques. But it hasn’t stopped there. PCR has greatly evolved over the years. Today, we stand at a point, where we can clone micro RNAs (miRNAs) in real time!
Due to miRNA size (about 18-21 nucleotides long) and varied expression levels, detection and quantification of miRNAs is a technically challenging task.
However, the development of the stem-loop real-time PCR technique by Chen et al.1, provides “…a novel real-time quantification method for accurate and sensitive detection of miRNAs and other small RNAs”.
Advantages to Stem-loop Real-time PCR
Overall, the stem-loop real-time PCR technique provides the following advantages:
- The technique is highly specific for miRNAs, and nonspecific genomic DNA is not amplified.
- The technique is specific to mature miRNAs and not their precursors, even if both are present in equal concentrations.
- You can use whole cell lysates as well as total purified RNA.
- Finally, you can start with a total RNA concentration as low as 20 picograms to quantify miRNAs.
The simplified scheme for stem-loop real-time PCR includes two very familiar steps: Reverse transcription (RT) and real-time PCR. But what makes it work with miRNAs is a stem-loop primer.
The Key: The Stem-loop Primer
The stem-loop primer is the main ingredient that makes miRNA amplification feasible. These RT primers are highly specific because of the way they are designed.
The stem-loop primer consists of a constant region that forms a stem loop and a variable six- nucleotide extension. The stem-loop portion of the primer extends the ~22 nucleotide miRNA to more than ~60 nucleotides to allow for traditional PCR in subsequent steps. The six-nucleotide extension is the reverse complement of the last 6 nucleotides on the 3’ end of the miRNA of interest and provides specificity. For a great picture of a stem-loop primer and the details of how it works, check out the original article.
The success of stem-loop primers over linear primers could possibly be attributed to the stacked bases in a stem-loop, which provide thermal stability and spatial constraint that minimize errors of primer binding.
For specific information on how to design your own stem-loop primers, you can visit miRNA Design tool.
Reverse Transcription and Real-time PCR
For amplification, the miRNA is reverse transcribed using the stem-loop primer. Next, an miRNA specific forward primer, a universal reverse primer (specific for the stem-loop portion of the primer) and a dual-labeled TaqMan probe is added to the reaction to initiate a typical real-time PCR.
Troubleshooting Stem-loop Real-time PCR
You should consider several factors in optimizing your protocol.
- The stem-loop primer needs to be properly folded. If it is unfolded, non-specific transcription may occur. Refolding of the primer into a stable structure takes place overnight in a regular thermocycler.
- Beware of rare miRNAs: An miRNA of interest that’s not abundantly expressed can potentially generate off-targets and nonspecific quantitation. However, a combination of specific forward primer-probe pairs should minimize this noise.
- Use proper controls: Make sure you include negative and positive controls. Test your protocol using synthetic RNA standards. It will help you dissect the reason for any nonspecific amplification/
- Be sure that you’re using the proper equipment (i.e., PCR hood, filter pipette tips, etc.).
- You need to know the precise sequence of the mature miRNA target, as this will determine the 3’ extension sequence of your stem- loop primer.
Stem-loop RT-PCR is a cost effective, convenient and consistent technique to quantify and clone miRNAs, as well as other small RNA. For once, cloning miRNAs won’t throw you for a loop!
Did you like this article? Make sure you read other PCR specific article, like ones to help you prevent PCR contamination.
References
- Chen C et al. (2005) Real-time quantification of microRNAs by stem–loop RT–PCR. Nuc. Acids Res. 33(20), e179.