Since 1993, MO BIO Laboratories has been developing innovative tools for researchers in Molecular Biology. MO BIO’s PowerMAX® DNA Isolation kits and our RNA PowerSoil® kits are among the most unique and innovative products in the biotech marketplace today. Effectively removing humic acid inhibitors of PCR present in soil samples. MO BIO’s line of soil and microbial isolation kits are now the method of choice among environmental and microbiology researchers studying microbial DNA in soil samples.
FFPE Tissues: How to isolate pure DNA without xylene
Content sponsored by MO BIO

Formalin fixed, paraffin embedded (FFPE) samples are derived from tissues (usually suspected tumor samples) that are fixed with formalin to preserve the cytoskeletal and protein structure and then embedded in a type of paraffin wax so the tissue can be sliced on a microtome, an instrument used to prepare very fine slices, 5-10 microns thick. Formalin irreversibly cross-links proteins via the amino groups, preserving the structural integrity of the cells so they can be stained with dyes used to analyze for abnormalities in the tissue that indicate cancer. However, the effect of these cross-linking fixatives on the nucleic acids is detrimental. Isolation of nucleic acids is impaired by both the paraffin wax and the cross-links that block DNA polymerases and inhibit PCR if they are not removed.
The outlook for getting DNA from these samples sounds pretty grim. Fortunately, we’ve overcome both of these major obstacles and can isolate pure, high quality DNA from FFPE tissues with no problem.
How do I remove paraffin wax from FFPE samples?
The traditional method for removing the wax has always been to use xylene, a highly flammable organic solvent. The tissue is washed several times in xylene to dissolve the wax and then the xylene is removed by performing multiple washes with ethanol before DNA isolation. This results in many extra handling steps where the tissue is repeatedly washed, and each time some of the tissue may become dispersed with the wax and removed with the solvent.
Another approach is the non-toxic BiOstic Paraffin Removal Reagent. This works very similarly to xylene but is safe and biodegradable.
If I skip wax removal, can I get higher yields of DNA?
Yes you can. It isn’t necessary to remove the wax if you have a method for increasing the activity of proteinase K to allow for digestion right through the wax. This reduces handling time and loss of tissue.
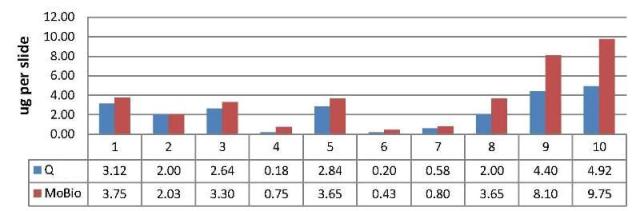
To achieve this effect, we’ve come up with a combination of solutions that results in a strong denaturing environment and promotes proteinase K activity at elevated temperatures resulting in complete digestion of the tissue while the wax is melting. If you’ve used the BiOstic FFPE Tissue DNA Isolation Kit, you know these as Solutions FP1 and FP2. The synergy between these two solutions surpasses the level of activity achieved using standard proteinase K digestion buffers, resulting in higher yields of DNA as demonstrated in Figure 1 below. In this example (provided by a customer) using 10 micron thick single slices removed from histology slides, the samples in blue were pre-processed with xylene and DNA isolated following the a protocol from manufacturer Q, and the MO BIO samples (red) were extracted directly without wax removal steps and DNA was purified using the BiOstic FFPE Tissue DNA Isolation Kit. Yields were quantified on the NanoDrop. As you can see, yields can range from 0.5 ug up to 10 ug per slice with an average of 1-2 ug for these samples. Not all slices are alike, but the BiOstic FFPE Tissue DNA Isolation Kit provided higher DNA yields in all cases.
Figure 1. DNA Yield of samples isolated using two methods.
Molecular weight of DNA isolated from FFPE tissue samples
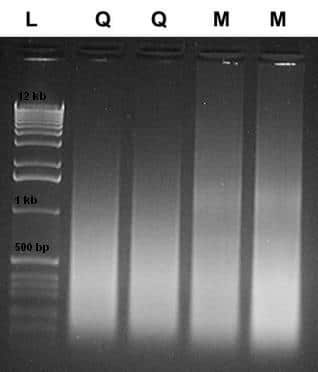
The size of DNA isolated will be mostly influenced by the fixation process and how long it was fixed. Size can also be affected by the age of the tissue sample. Typical sizes are generally small, around 100-500 bp (see Figure 2 for example). Here, samples were isolated using the kit from manufacturer Q, as described above (lanes labeled Q), or the BiOstic FFPE Tissue DNA Isolation Kit (lanes labeled M). Using the BiOstic FFPE Tissue DNA Isolation Kit, which avoids solvents and extra handling, may prevent further DNA damage and loss (note the higher molecular weight DNA in the “M” lanes, processed without xylene). You really don’t know what your sample will look like until you’re done, therefore we always recommend using a qPCR assay designed for the smallest possible amplicon.
Figure 2. Agarose gel analysis of DNA isolated using two methods.
Obviously, degradation is pretty severe for these sample types. They aren’t handled in an RNase-free environment and the tissues can be many years old, sitting on slides in drawers at room temperature. But people are isolating RNA from FFPE tissues and generating informative data, so it can be done.
RNA isolation is possible using the BiOstic FFPE Tissue DNA Isolation Kit as well, with some minor protocol changes. The FP1 and FP2 combination is too harsh for RNA so we switched this out for another buffer with RNA protective qualities and a neutral pH (called BF2). Additionally, the temperature for removing the protein cross-links can’t be performed at 90degrees Celsius like it is for DNA. In the paper by Masuda et. al., Analysis of chemical modification of RNA from formalin-fixed samples and optimization of molecular biology applications for such samples, Nucleic Acids Research, 1999, vol. 27, No. 22, pages 4436-4443, a temperature gradient was used to determine which temperature and for how long can the RNA be incubated to remove the protein cross-links and result in successful RT-PCR. They found that 30 minutes at 70 degrees Celsius was sufficient to do the job.
Contact MO BIO technical support by emailing technical@mobio.com to enquire about our modified protocol for RNA isolation from FFPE tissue. You can request a free sample of the BiOstic FFPE Tissue DNA Isolation Kit here.
Isolation of nucleic acids from FFPE tissues made easy!
FFPE Tissues are a unique sample type with a lot of challenges, but when it comes to DNA isolation, we’ve made that part easy. Samples are available and can be ordered on the web at no charge or upon request if you call technical support or customer service. We’re waiting to help you out!
