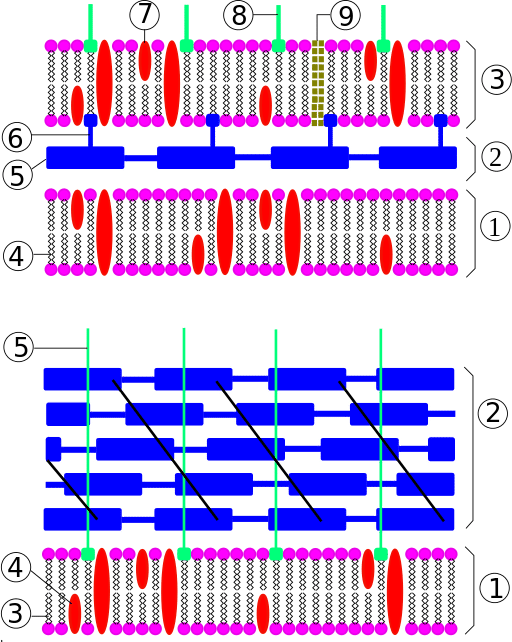
What is Adeno-Associated Virus?
Adeno-associated virus (AAV) is a popular gene therapy vector. Different AAV serotypes infect different cell and tissue types with varying efficiency, so it’s a good idea to select the serotype most relevant to your work. The most commonly used AAV serotypes are 1, 2, 5, 6, 8, 9 and DJ, although many other naturally occurring and recombinant forms exist.
Why AAV?
Unlike lentivirus, AAV very rarely integrates into the cell genome. This can limit its usefulness for experiments in rapidly dividing cultured cells, because the virus is rapidly diluted within the culture and may also be actively degraded. However, a non-permanent transduction may be useful for some applications, such as CRISPR. AAV is also an excellent choice in vivo, with various serotypes offering high transduction efficiencies in nervous tissues, lung, muscle and liver.
Before You Begin Working with AAV
Similar to lentivirus, AAV production begins with cell transfection. You will need the AAV helper plasmid, which contains the Adenovirus sequences necessary for AAV production. This helper plasmid is commercially available as pHelper or as pladeno5. Serotype plasmids are available from various sources. Keep in mind that many serotype plasmids are under a materials transfer agreement (MTA), so make sure you source and acknowledge them correctly! Finally, you will need your transfer plasmid containing your gene of interest and the AAV2 inverted terminal repeats (ITRs).
TIP: you don’t need serotype-specific AAV transfer plasmids. Production of most AAV serotypes utilises the AAV2 ITRs.
The two most common AAV transfer plasmids are the pTRUF series (available from the University of North Carolina vector core) and pAAV-MCS, available from Cell Biolabs. My preferred option is pAAV-MCS, as I have found it to be slightly more stable than pTRUF with more space for transgene sequences.
TIP: AAV transfer plasmids have SmaI sites in their ITR sequences. You can therefore cut the plasmid with SmaI if you need to check the integrity or size of your plasmid.
Lentivirus does not have a packaging ‘limit,’ as such. Lentivirus packaging efficiency predictably decreases as sequence size increases. In contrast, AAV has a packaging limit of 4.7kb. You can try to package larger sizes, but virus production efficiency will be markedly decreased. This 4.7kb of sequence must include the ITRs, promoter, your transgene, a marker (if you are using one), and a polyA sequence.
TIP: Minimal promoters and synthetic minimal polyA sequences can help to reduce packaging size, as can the use of 2A peptides over larger sequences such as internal ribosome entry site (IRES).
Start Here
The first step of your AAV production is cell transfection, and the transfection method you choose is up to you. If you’re using calcium orthophosphate (CaPO4), your cells should be about 40% confluent at the time of transfection. If you’re using a commercial lipid-based method or polyethyleneimine (PEI), they should be 90-95% confluent. I have found that CaPO4 produces higher AAV titres, but lipid based methods will still work fine. Bear in mind that you don’t want to use a transfection reagent that results in widespread cell death—you will be harvesting AAV from the cells. So, if the cells lyse, then AAV ends up in the culture media where it will be harder to recover. If you are using CaPO4 for transfection, it’s best to change the media 6-18 h after transfection to prevent cell death—I find that 6-8 h is best.
TIP: For CaPO4 transfection, there is no need to pH multiple HBS solutions to find the best one. Just add HEPES to your culture media before you transfect.
Harvest
Twenty-four hours post transfection, check your cells using fluorescence microscopy to make sure that the fluorescent reporter (if you’re using one) is present. At 72 h post-transfection, the AAV is ready to harvest. Virus particles will be present in both the cells and the culture media, but harvesting AAV from the cells is much easier. To harvest your AAV, follow the steps below:
- Remove culture media and set aside in a 50 mL tube. Add cold PBS supplemented with 10 mM EDTA to the cell monolayer. Wait 5 minutes, detach the cells and then add them to the 50 mL tube. Spin at 1500 rpm for 15 minutes, and discard the supernatant.
- Resuspend your cell pellet in 800 mL of AAV freezing media (see footnote for recipe). Freeze thaw 3 times—incubate at -80°C for 20 minutes and then 37°C for 5 minutes
- Briefly spin. Then, add Benzonase enzyme to a final concentration of 50U/mL. Incubate at 37°C for 30 minutes.
- Spin in a microcentrifuge at 14,000 rpm, 4°C for 30 minutes. Remove and keep supernatant.
This is your crude AAV prep. If you’re using a plasmid expressing an RFP or GFP, this lysate will often be bright green or red.
TIP: If you want to test your AAV on cultured cells, this lysate is perfectly fine to use as is—further processing of the AAV will increase the purity of the prep but not necessarily the concentration. Crude AAV should not however be used in vivo or on very sensitive primary cells because of various cellular proteins that will still be present in the lysate. If you aim to use a fluorescent plate reader, the presence of soluble fluorescent proteins can also produce very high background readings.
AAV Freezing Buffer Recipe:
- 4 mL H2O
- 1 mL 1 M Tris pH 8
- 5 mL 5 M NaCl
- 1 m 1 M MgCl2
The crude AAV can be stored at -80°C until needed. If you want to freeze the lysate and purify it at a later date, then that will work perfectly fine. For the next stages of AAV purification, keep a lookout for our next article in this series, AAV Production Part II: Purification and Transduction.