Bitesize Bio: Life Science & Bioscience Articles for Researchers
Latest Events
Latest Articles
Performing Pipette Calibration Yourself
It’s necessary to check pipette calibration every few months to ensure accuracy by dispensing the right volumes. This article explains how to perform pipette calibration yourself so you can ensure they are accurate and avoid the wait for their next service.
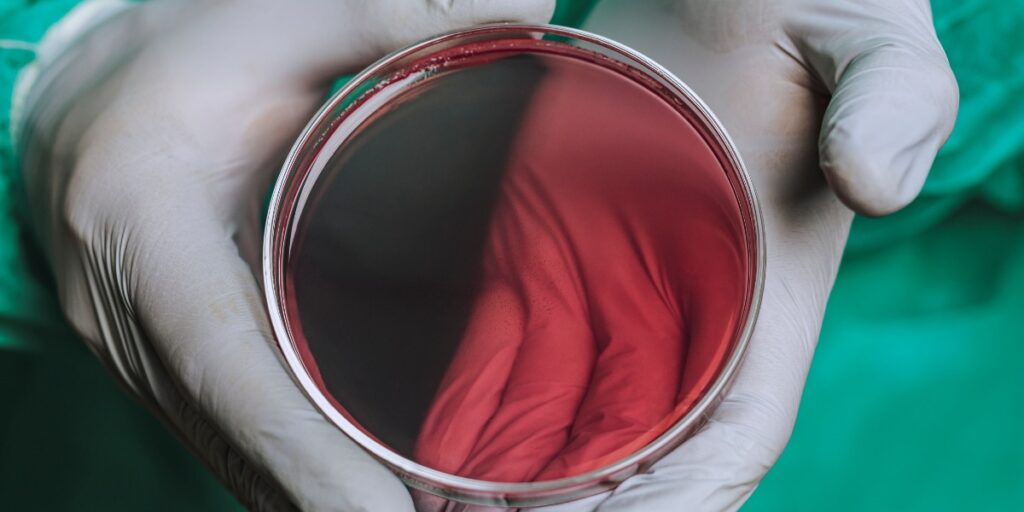
Agar plates are the foundation of many experiments. Make sure your plates are perfect every time with these 8 tips and start your experiment ready for success!
Nucleic acid extraction kits are routinely used in today’s molecular biology labs. Read on to learn more about what is inside these black boxes of wonder and how you can get the best results for your preps.
Biomarkers are fundamental in bioscience, from basic microbiological experiments to clinical studies. This article explains the different types of biomarkers and their application in cancer and disease research.
Do you hate it when you get into the lab ready to get stuff done and then find yourself spending the first 45 minutes of the day hunting for what you need? We do too. So read these 10 tips to organize your lab space for better productivity and spend less time hunting for stuff and more time doing research.
Microscopy & Imaging
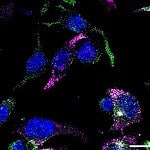
Adherent cell fixation is a crucial step in preparing cells for microscopy and imaging, ensuring that cellular structures are preserved for detailed analysis. Read our 8-step guide on how to effectively fix adherent cells to your microscope slides, including tips on sterilization, coating, and fixation methods, right here.
DNA / RNA Manipulation & Analysis
Nucleic acid extraction kits are routinely used in today’s molecular biology labs. Read on to learn more about what is inside these black boxes of wonder and how you can get the best results for your preps.
How-To Guides
Article Categories
- Analytical Chemistry & Chromatography Techniques
- Cells & Model Organisms
- Chemistry for Biologists
- Cloning & Expression
- DNA / RNA Manipulation & Analysis
- Flow Cytometry
- Genomics & Epigenetics
- Microscopy & Imaging
- PCR, qPCR & qRT-PCR
- Bioscience Mastery
- Basic Lab Skills & Know-how
- Career Development & Networking
- Dealing with Fellow Scientists
- Equipment Mastery & Hacks
- Getting Funded
- Lab Safety
- Lab Statistics & Math
- Organization & Productivity