A Crash Course in Epigenetics Part 1: An intro to epigenetics

These days, epigenetics is a fast moving field. I don’t remember having learnt about it during my biomedical studies, some 10 years ago. Nowadays, there seems to be no way around it when studying health and disease. Increasing interest combined with recent technological breakthroughs have led to quickly expanding knowledge of its abundant and important role in biology.
You probably think regulation of gene expression is complicated enough, with its transcription factors, enhancers, repressors, and post-transcriptional and post-translational modifications. But epigenetics adds yet another layer of complexity and it can be a bit overwhelming if you have not kept pace with its development.
So in this series of 4 bite-sized articles I will give you an easy-to-digest crash course in Epigenetics; What it is, the tools used to study it, how it is regulated and the potential therapeutic targets.
Grab a coffee and let’s get started…
A quick definition
Epigenetics refers to heritable changes in gene expression caused by non-genetic mechanisms, thus by alterations other than in the DNA sequence. Epigenetic changes can persist throughout an organism’s lifetime and be passed on to multiple generations.
A little bit of history
In 1942, Sir Conrad Waddington pondered how environmentally triggered traits could become inherited features of an organism. In Nature, he published his theory of genetic assimilation, hypothesising that organisms are normally protected against minor environmental and genetic variations. The protective buffering capacity might be reduced under extreme stress, he thought, allowing expression of previously latent genetic variation. Repeated rounds of selection through continuing stress would allow the adaptive phenotype to persist, to ultimately become independent from the original environmental stimulus.
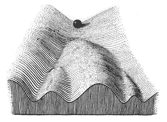
Although experimental tools were limited at the time, in 1953 Waddington demonstrated the existence of genetic assimilation of an acquired character under environmental stress, in Drosophila. These and other observations led him to propose his now famous epigenetic landscape, which laid out the basis for epigenetics as we know it today. In this landscape, he imagined marbles rolling down a hill, competing for a passage through different grooves on the slope. The marbles come to a halt at the lowest point, representing cell fate. Mutations alter the grooves of the slope, and thereby the cellular developmental process, ultimately affecting cell differentiation. Although he thus proposed an interaction between the environment and genetic information, he had no idea about the underlying mechanisms. Not until the 1990s did a renewed interest in genetic assimilation lead to elucidation of the molecular basis of Waddington’s observations. Since then, much effort has been put into unraveling epigenetic mechanisms.
 I will discuss different components of the epigenetic machinery and how scientists have discovered their existence, as well as shed light on their functioning during development, in health and disease, and how they might prove to be attractive therapeutic targets.
I will discuss different components of the epigenetic machinery and how scientists have discovered their existence, as well as shed light on their functioning during development, in health and disease, and how they might prove to be attractive therapeutic targets.
Chromatin: where it all happens
In order to understand how epigenetic mechanisms function, we first need to appreciate chromatin. Chromatin is the ordered structure of genomic DNA and associated proteins in the nucleus. It is the natural state of DNA in the nucleus: all DNA-related processes function in the context of chromatin, rather than on naked DNA. The highly organised manner in which DNA is wrapped around specialised histone proteins makes it possible to package 2 meters of DNA inside a cell nucleus with a diameter of ~6 um, while at the same time allowing it to easily become accessible to a variety of proteins involved in DNA transcription, replication and repair processes.
The epigenetic machinery
A complex of eight histone proteins, and a stretch of double-stranded DNA 146 nucleotides long that is wrapped around the histone core, form a nucleosome. Different types of histone proteins exist; a nucleosome is made up of two copies of each of the core histones H2A, H2B, H3 and H4. They have a protruding N-terminal amino acid ‘tail’ in common. These histone tails are subject to a wide variety of modifications, which form a central component of epigenetic gene regulation. Amino acid residues in histone tails can for instance be acetylated (lysines), methylated (lysines) or phosphorylated (serines). Acetylation is generally associated with active transcription, but for most other modifications the effects are less predictable.
Besides histone modifications, epigenetic mechanisms include chromatin remodelling and nucleosome repositioning, DNA CpG methylation, and some would include the effect of small and non-coding RNA-mediated regulation. Chromatin can adopt a more or less condensed state. Chromatin remodelling complexes are responsible for changing the level of compaction temporarily. Loosening up the chromatin allows access to nucleosomal DNA, thus facilitating DNA processes such as transcription, replication and repair. Histone modifying enzymes and chromatin remodelling complexes work in concert to alter chromatin conformation. Certain histone modifications attract specific remodelling complexes.
At the same time, DNA methylation may recruit methyl-binding proteins, which form a complex with histone modifying enzymes. Thus, the net effect on gene transcription is the result of the combination of different histone modifications on adjacent residues of histone tails, as well as of DNA methylation and associated proteins and the action of chromatin remodelling complexes. All these processes together yield a remarkable cellular diversity, with identical sequences as its basis in every cell.
The (slightly) bigger picture: epigenetic processes in a cellular context
Impressive high-throughput technologies that will be touched on in the next article in this series have provided linear maps of epigenetic marks. Although these maps are very insightful, keep in mind that the next logical step would be to move on to 3D representations that link epigenetic processes to cell signalling cascades and environmental clues. Although epigenetic mechanisms take place in the nucleus, they can occur in response to environmental signals, such as hormones, nutrients, stress and cell damage. This indicates that extracellular and cytoplasmic factors are also at stake in epigenetic regulation.
How exactly non-epigenetic cues induce cells to alter their epigenomes is an important question that needs answering. How is it possible that genes are generally under stringent epigenetic control, and only get activated and transcribed when needed?
Certain cellular signalling pathways have already been earmarked as candidate regulators of epigenetic remodelling. Not surprisingly, they tend to be factors with already established roles in cell lineage commitment, growth and development.
Then what makes epigenetic regulation distinct from classic signal transduction? The latter refers to direct changes in gene expression due to altered levels, post-translational modifications, or actions of transcription factors, in response to the extracellular environment or state of the cell. Once gene transcription has been changed, a new epigenetic signature might be established to stabilise the signal transduction, thereby rendering it heritable and independent of the initial triggering signal.
Of course, that is not all: as enzymes of the epigenetic machinery can be regulated by signalling molecules, but might at the same time affect the activity of signalling molecules, the interplay between classic regulatory signals and epigenetic control is – you guessed it – complex and likely often intertwined. Indeed, certain signal transduction effectors such as transcription factors and signalling molecules have been found to interact with chromatin modifying enzymes.
In the next article in this series, I will discuss some experimental techniques that have helped us better understand these processes.
Sources/further reading:
Portela A, Esteller M (2010) Epigenetic modifications and human disease. Nat Biotechnol 28: 1057-1068.
Mohammad HP, Baylin SB (2010) Linking cell signaling and the epigenetic machinery. Nat Biotechnol 28: 1033-1038.
More in this series:
Part 1: An introduction to Epigenetics
Part 2: The Toolbox of the Epigeneticist
Part 3: Regulated Regulation
Part 4: Disease Mechanisms and Therapeutic Targets
1 Comments
Leave a Comment
You must be logged in to post a comment.
I am interested in epigenetics are there any courses