Basics of Protein Phosphorylation Part III: Family Ties – Diversity and Similarity Among Protein Kinases
So far in our “Basics” series, we’ve taken an overview of the major players involved in protein phosphorylation, and some of the tools that one might use to study them. Now, we’ll return to the protein kinases to take a deeper look- though our “deeper” look will barely scratch the surface, as there are currently over 500 known kinases with several thousand targets. We’ll talk about the amino acids phosphorylated by kinases, the families of protein kinases, and how they choose their targets.
Three’s a crowd
20 amino acids are shared amongst all forms of life, used to build the proteins that allow the biochemistry to take place. Out of those 20, several have the electron-rich functional groups containing oxygen, nitrogen, and even sulphur to which an inorganic phosphate group could be added. Even so, only three amino acids are most commonly recognized as phosphorylatable: serine, threonine, and tyrosine. Kinases generally fall into two categories: they will either phosphorylate the smaller serine and/or threonine- usually interchangeably- or they will phosphorylate the bulkier aromatic tyrosine. They are thus appropriately named the protein serine/threonine kinases (PSTKs) and protein tyrosine kinases (PTKs). As easy as it would be to say that these are strict barriers- especially in a “Basics” article- nothing in biosciences is ever so rigid. Some kinases have the ability to phosphorylate both serine/threonine and tyrosine. Those that are well-known to do so are termed dual-specificity kinases, or mixed kinases.
While these 3 phosphorylated amino acids exist in eukaryotic proteins, other amino acids can, less commonly, be phosphorylated. Histidine kinases, for instance, are more prominent in bacteria and barely seen in humans. That said, each passing year allows more and more histidine kinase discoveries to creep into mammalian systems.
The code (and no… not Da Vinci’s)
All proteins have at least one serine or threonine present in their primary amino acid sequence. Realistically, they have many, many more than one. Does that mean that a PSTK will phosphorylate every single serine and threonine in every single protein? Clearly, they won’t. Just as all enzymes have a discrimination and recognition for their appropriate substrate, so too do kinases have recognition for which sites they should be phosphorylating. In the process of phosphorylating an amino acid, kinases also recognize and bind to adjacent amino acids towards both the N- and C-termini of the phosphorylation site, known as a consensus sequence. The presence or absence of these consensus sequences determine whether a certain protein will be phosphorylated by a given kinase. Recall even from my previous article on tools that phospho-specific antibodies- another biochemical interface like a kinase binding site- will also recognize not only a phosphorylated amino acid, but also an “epitope” of adjacent amino acids.
Consensus sequences depend on certain characteristics of select amino acids- polar, nonpolar, acidic, basic- while allowing for other positions to be flexible. This gives kinases their ability to have more than one substrate. Consider, for example, the following amino acid sequence:
Protein sequence: SEPIPLESREEYMNG
The blue S is the serine to be phosphorylated. Now, compare this to the consensus sequence for casein kinase 2 (CK2), an acidophilic PSTK recognizing sequences generally containing negatively-charged amino acids, like aspartate and glutamate.
Protein sequence: SEPIPLESREEYMNG
CK2 consensus: [S/T]xx[E/D]
According to this consensus sequence, CK2 will phosphorylate a serine/threonine which has an acidic amino acid, either aspartate or glutamate, at the third position towards the C-terminus. The two amino acids in-between are entirely flexible. It is this kind of flexibility, and the combinations of amino acids that could fit into those two in-between positions that allows CK2 and other kinases to have many target proteins. Now, consider another sequence:
Protein sequence: QHRARQKTLEHLQLS
Protein kinase B (PKB, or more commonly known as Akt), a basophilic PSTK, will phosphorylate sites with regions of positively-charged amino acids. Compare its consensus sequence:
Target sequence: QHRARQKTLEHLQLS
Akt consensus: RxRxx[S/T]
While three amino acids in this consensus sequence can be flexible, it is required that the third and fifth amino acids to the N-terminal of the phosphorylation site be arginines.
All in the Family
The amino acid phosphorylated, and the consensus sequence at which a kinase will phosphorylate, are two important characteristics. When you take these two characteristics together and categorize them, you begin to form groups of kinases- or as they are more appropriately called: families.
Cell Signaling Technology has a very nice diagram of the family tree of the kinome on their website. It’s one of my personal favourites and a big help. Let’s take a moment to go over the families listed:
AGC family: Named after some of the best-known, textbook kinases- protein kinase A (PKA), protein kinase G (PKG, and protein kinase C (PKC). Akt/PKB is also present in this family. Many of these PSTKs tend to fall in the insulin signalling pathway.
CAMK family: Named after the Ca2+/calmodulin-dependent protein kinase (CaMK), activated by calcium signals. Also included in this family is AMP-dependent protein kinase, another “sensor” kinase which is activated upon depletion of energy stores and accumulation of AMP. Additionally present are LKB1 (which activates AMPK), and the mitogen-activated protein kinase-activated protein kinases (MAPKAPKs).
CMGC family: This family includes the cyclin dependent kinases (CDKs). Additionally present are the c-Jun N-terminal kinases (JNKs), the extracellular signal-regulated kinases (ERKs), the p38 isoforms, which are all considered members of the subfamily of mitogen-activated protein kinases (MAPKs). Also includes glycogen synthase kinase-3 (GSK3).
GYC family: Arguably, the most important member of this family is casein kinase 2 (CK2), which presently has over 200 known substrates, including other protein kinases in cell cycle, DNA repair, oncogenesis, and insulin signalling. Also present is aurora kinase, which regulates mitotic spindle dynamics, and inhibitor kappa B protein kinase (IKK), which mediates inflammation.
TK family: The tyrosine kinase family. Many of these are receptor tyrosine kinases (RTKs), linked to receptors such as the insulin receptor, epidermal growth factor (EGF), platelet-derived growth factor (PDGF), and so forth. The RTKs are the highest protein kinases in signal transduction hierarchy. Also includes non-receptor tyrosine kinases like the Janus kinases (JAKs)
TKL family: This family contains an eclectic assortment of PTKs involved in a multitude of biological processes. The interleukin receptor-associated kinases (IRAKs) are important in cytokine and chemokine signalling. Others, like Raf1, are downstream of the Ras family, which are associated to G-proteins and involved in activating MAPK pathways.
STE family: The sterile 20-like family includes PTSKs that regulate the members of the CMGC family. Whereas the CMGC family includes MAPKs, the STE family includes the MAPKKs (MAP2Ks) and the MAPKKKs (MAP3Ks).
CK1 family: Similarly-named, but apparently not related to CK2. A very small family, it essentially only includes the various isozymes of CK1, as well as the tau tubulin kinases (TTBKs) and vaccinia-related kinases (VRKs).
An intricately-woven tapestry
As I’ve pointed out throughout this series, there are various ways in which you can study protein phosphorylation. You can study a specific target, a specific protein kinase, a pathway- really there’s a vast spectrum. What I hope this article helps to illuminate for Bitesize Bio readers is that, in the world of protein phosphorylation, it’s very difficult to stay on one narrow theme and remain fixed on that alone.
For example, you might be interested in glycogen metabolism, and how it’s polymerized by glycogen synthase, and broken down by glycogen phorphosylase. Ultimately, though, it’ll become hard to ignore the concepts of phosphorylation and activation of glycogen phosphorylase by phosphorylase kinase and PKA of the ACG family, and the concurrent phosphorylation and inactivation of glycogen synthase by GSK-3 of the CMGC family and by PKA of the ACG family. GSK-3, in turn, can be phosphorylated and inactivated by Akt of the ACG family upon insulin stimulation, allowing glycogen synthase to remain active. Dephosphorylation of glycogen synthase by protein phosphatase 1 (PP1) also activates it.
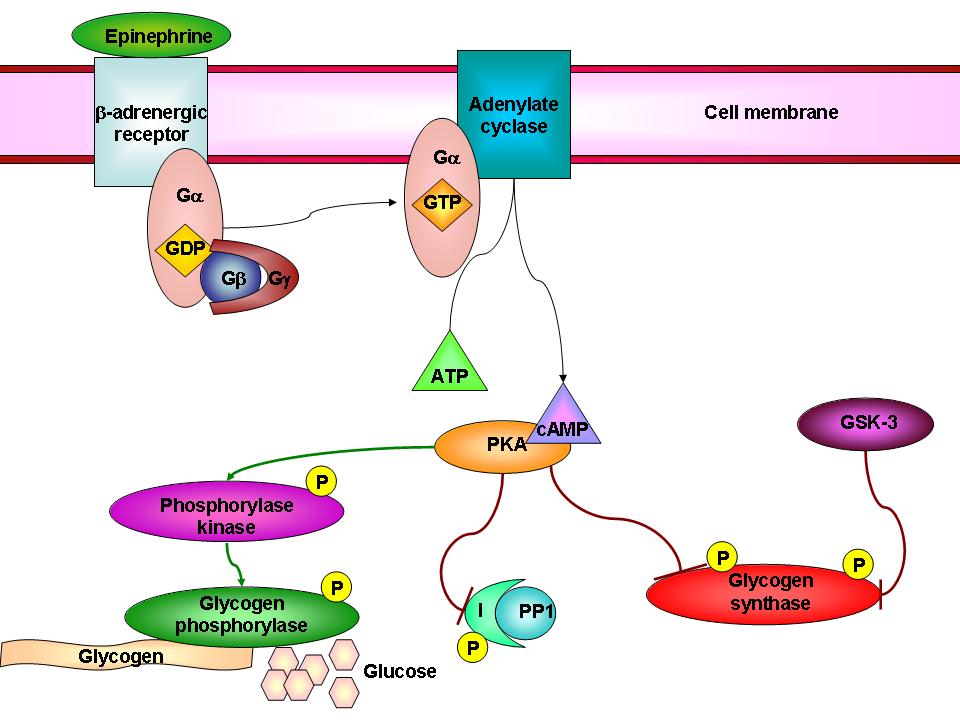
And thus, in this small example of two metabolic enzyme, three kinases and one phosphatase, we see how complex the story can be- but more importantly how integral protein kinases (and phosphatases) are to metabolic regulation.
Next, to cap off the “basics” of protein phosphorylation, we’ll take a look at the enzymes that can so easily tear down the complex structure built by protein kinases: protein phosphatases.
6 Comments
Leave a Comment
You must be logged in to post a comment.
Hello Christopher,
This content is very helpful. Thanks very much.
And I notice there was a diagram about “Dephosphorylation of glycogen synthase” not showing. May I know if it can be replaced by this image?
https://ars.els-cdn.com/content/image/3-s2.0-B012227055X005630-gr1.jpg?_